How Tafamidis was born
 Read with caution! This post was written during early stages of trying to understand a complex scientific problem, and we didn't get everything right. The original author no longer endorses the content of this post. It is being left online for historical reasons, but read at your own risk. |
Months ago, I heard for the first time a remarkable story of how one family’s resistance to familial amyloid polyneuropathy (FAP) led researchers to discover a drug to treat that disease. This morning I finally got the full story. Jeffery W. Kelly of the Scripps Institute gave a lecture at the Hereditary Disease Foundation’s HD2012 conference recounting the history of how tafamidis was developed to treat FAP.
FAP is caused by aggregation of transthyretin (TTR), a protein that forms tetramers which transport retinol. Interestingly, in rodents it also transports thyroid hormone, and in humans the tetramer still has its thyroxin site, but this site is left empty in 99+% of human TTR tetramers simply because we also have two other, higher-affinity, thyroid hormone transporters which rodents lack.
The genetic disease FAP is caused by a V30M mutation in the TTR gene. FAP is an “in trans gain-of-proteotoxicity disease”: aggregated TTR destroys tissues other than the tissues that make it. Specifically, TTR is synthesized in the liver but its toxicity affects the heart and the autonomic nervous system.
By the early 1990s, Kelly’s group began to believe that the disease was caused by a dissociation of the TTR tetramer. Once a monomer is formed, they believed, there is a “downhill” (energetically favorable) path to aggregate formation, so the most viable place to intervene and prevent disease would be to stabilize the tetramer itself. Specifically, the group believed that the unstable point in the tetrameric structure is at the dimer-dimer interface– right where two thyroxin binding sites exist which are usually empty in humans. Interestingly, while mutant TTR causes FAP, wild-type TTR can also dissociate and cause senile systemic amyloidosis (SSA), which affects 15% of people over age 85 and leads to heart failure (read: “dying of old age”).
Based on what they were finding in the lab, the group had already begun to focus on the dimer-dimer interface as the potential road to therapeutic development. However, in 1993 their hypothesis got a major boost from one family’s genetic data. Coelho 1993, in an article entitled “A strikingly benign evolution of FAP in an individual found to be a compound heterozygote for two TTR mutations: TTR MET 30 and TTR MET 119.” (J Rheumatol 20, 179) identified a family in Portugal which carried the V30M mutation yet had never shown signs of FAP. Sequence data revealed that on this family’s other allele, they had a polymorphism T119M which protected them from the disease. In Kelly’s words, the conclusion they drew from this finding was that “T119M TTR protects in trans through mixed tetramer formation”. In other words, the activation energy required for a mixed tetramer (with the codon 119 heterozygosity) to dissociate and form amyloid is significantly raised. Kelly’s group published two Science papers documenting the effects of this mixed tetramer: Hammartstrom 2001 did a bunch of biochemistry to determine the rate of tetramer dissociation and amyloid deposition in “mixed” (some T119M) tetramers versus all-119T tetramers, and Hammarstrom 2003 showed that small molecules were able to interfere with dissociation in the same way that the T119M polymorphism does.
Kelly describes the thinking as follows: we knew we needed a small molecule that would bind to this binding site at the dimer-dimer interface, but that was neither an agonist nor antagonist for thyroid hormone itself. The chemists set out to design such a molecule and seem to have struck gold by inventing Tafamidis. It was first developed in or around 2003 and it has already completed a 30-month clinical trial with 100 or so participants, and was approved in the E.U. (under the name Vyndaqel) in November 2011. After reviewing Tafamidis this year, FDA still wants a second clinical trial. Still, it’s a pretty fast progression for a drug, and the data that Kelly showed on the case vs. control groups was remarkable– not only did Tafamidis perform much better than placebo, it also reversed some of the decline in people who switched from placebo to Tafamidis after the 30-month trial ended. Kelly also believes Tafamidis will prove useful in treating SSA, which would be a much larger market. Interestingly, he also pointed out a study in Denmark (I believe it was Hornstrup – A stabilizing variant in transthyretin; but I can’t find it online, maybe it is still in press) which, though the sample size was only about 50 people, suggested that people (not FAP sufferers, just any random people) with the T119M polymorphism live a few years longer than others, particularly among people who suffer heart attacks. About 1 in 200 people, at least in Denmark, carry the T119M polymorphism.
This whole remarkable story provides a compelling link between genetics and therapeutic development. It’s a strong motivation for the use of genome-wide association studies to find variants that protect people from genetic diseases they’re otherwise carriers for, whether by delaying onset or reducing severity. And it also highlights the value of having people at risk for genetic diseases contribute their genomes to science. I heard at HD2012 yesterday that something like 95% of people from HD families never get tested for HD. Much of this is because people figure if the disease is untreatable anyway, there’s no benefit to knowing, so it’s easier not knowing, less painful to just leave it at 50/50. But science needs as many genomes as it can get its hands on, because if we discover an individual with expanded CAG repeats in HTT or a D178N mutation in PRNP but is mysteriously asymptomatic, maybe we can unearth a genetic cause for this comparable to the T119M for FAP, that would help lead us to a therapeutic. Luckily, the two viewpoints are not mutually exclusive. It is possible for people to contribute their genetic data to science on the condition that they themselves never learn whether they carry the disease, and that the data be strictly limited to scientists working on the genetics of the disease. Indeed, I spoke with someone at the CJD Surveillance center at Case Western who specifically stated they would support such an arrangement on request. I would bet most, if not all, clinical outlets for genetic disease testing would offer the same. This solution respects people’s choice to not know their test results while also helping to advance the science and lead us toward a cure.
After hearing Kelly’s talk, I had some brainstorms. We actually already know of one polymorphism which seems to confer resistance to prion disease: the heterozygous G127V substitution that renders some Fore people resistant to kuru. If we could figure out how this prevents prion formation at the molecular level, could that be a basis for designing a small molecule to do the same? Because kuru is variant, this has never been tested in a genetic model. Perhaps a first step would be to create knock-in mice with haplotypes 127G 178N and 127V 178D and see if the mice get FFI symptoms. I sidled up to Kelly afterwards to ask him his thoughts. He was a super nice guy and supportive even if a bit skeptical of my idea. He said you could do that with mice and it might be interesting, but he cautioned against putting too much stock in the parallel between PrP and TTR. TTR is natively a tetramer, so stabilization of the tetrameric state was a natural goal for a small molecule. PrP by contast is natively a GPI-linked monomer, so at what stage in the cascade towards aggregation would you intervene? Would you try to halt it at the dimer stage? The tetramer stage? And if you did, would that halt the disease?
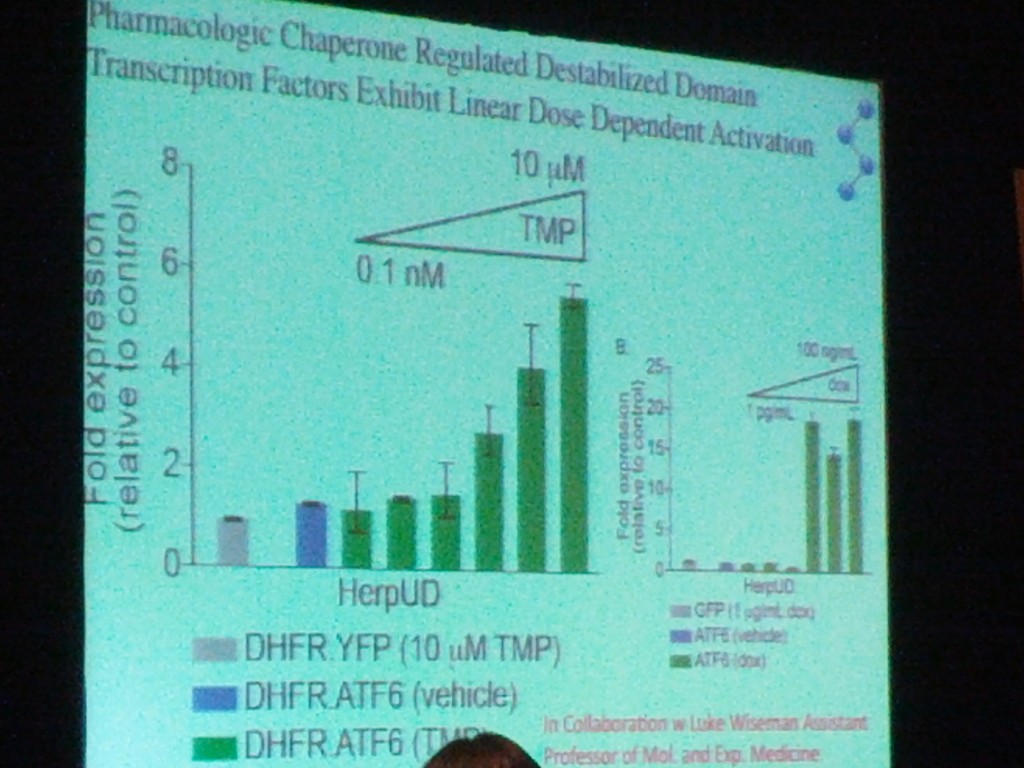
Another really important difference between TTR and PrP is that TTR’s native state is highly ordered, while PrP is what Kelly calls an “intrinsically disordered protein” — he also puts α-synuclein, Aβ, and HTT exon 1 into this category. This means that these proteins don’t just have a single stable “native” state. Instead, they adopt many energetically similar states in order to interact with many different molecules. (For PrP I think this is especially true of the unstructured N-terminal tail). A theme of Kelly’s talk was that for these intrinsically disordered proteins, it will be somewhat tougher to design a stabilizing small molecule akin to Tafamidis; instead, the approach he is more optimistic about is designing molecules that ratchet up the action of the proteostasis network — the multitude of different pathways encompassing 2500 or so proteins, many very highly conserved, responsible for maintaining protein homeostasis in each cellular compartment (the cytosol, the secretory pathway, mitochondria, etc.) (In fact, the proteostasis network is actually what Kelly’s talk was primarily about; the history of how Tafamidis was developed that I’ve documented here was more of a sidebar). Kelly is experimenting with molecules that can intervene in these pathways, for example, to direct more of a given protein towards degredation rather than secretion, thus reducing its concentration in the extracellular environment. He touched several times on the fact that aggregation is a concentration-dependent process, so reducing the concentration of HTT, PrP, etc. could slow disease even if the mutant form is still present in some concentration. (This makes perfect sense with Jackson 2009‘s finding that he could induce prion infection by intracranial injection of FFI brain homogenate only in mice that overexpress PrP.) In fact, Kelly also mentioned that doxycycline is involved in promoting degradation and reducing protein concentration, a fact I was unaware of. Now I need to go back and read more about the doxycycline studies that led up to the current clinical trial. Kelly showed this slide, which didn’t come out very clear in the photo but I think is supposed to demonstrate how the relationship between dosage and fold expression is more continuous for TMP than for doxycycline (the triangle at right is labeled “dox”):
Interfering in the proteostasis network sounds like a really promising road to therapeutic development for all neurodegenerative diseases. While Kelly’s group is already having some success with this approach, he was quick to point out we need to understand a whole lot more of the biology of these pathways in order to know how to effectively intervene.