Resources for systems biology in prion disease
 Read with caution! This post was written during early stages of trying to understand a complex scientific problem, and we didn't get everything right. The original author no longer endorses the content of this post. It is being left online for historical reasons, but read at your own risk. |
Systems biology is the science of using large datasets to infer what molecular interactions are happening and what biological networks are getting disturbed at the root of the disease or biological phenomenon you’re studying. When I first studied bioinformatics, my instructor told us a story about a drunk looking for his lost keys under a street light – not because that’s where he had lost them, but just because that was the only place with good enough light to bother looking for anything. This was an analogy for much of what we do in science. We’d love to watch PrP interact with the proteins it interacts with, see exactly what happens to convert it to PrPSc, and witness what that isoform does to cause trouble in the cell. But since everything in biology is too small to see, we look where the light is better.
The good news is that there are a lot of bright lights these days. In particular, arrays and sequencing have made everything to do with nucleic acids pretty high throughput: genotypes, expression levels, epigenetic markers, protein-DNA interactions. These things aren’t always what we actually want to measure: for instance, in many cases, protein levels would be much more informative than RNA levels, but proteomics is still much more expensive than expression arrays or RNA-seq. But they’re cheap enough to let us generate heaps of data that we can sift through to look for hints as to what’s really going on.
Hwang 2009 provides us with a landmark demonstration of this systems approach applied to prion disease. At the core is the use of microarrays to measure gene expression levels in brain tissue of prion-infected and control mice and thus determine which genes are expressed at higher or lower levels as a result of prion disease. Importantly, Hwang varied the prion strain (RML, 301V), time of measurement (60 to 350 days post infection), and the mouse strain: three unmodified inbred lines - B6, B6.I, FVB and three genetically modified lines – Tg4053 (which overexpress PrP 8-fold), heterozygous and homozygous PrP knockout mice. The large number of different combinations made it possible not only to cancel out biological and technical noise but also to examine which effects were prion strain-specific and which were shared, and which effects were time-dependent (or PrPSc concentration-dependent). In all, 333 genes were found to be differentially expressed in all combinations tested. These were then assembled – using a wealth of publicly available annotations on function, cell-specific expression, interactomics, and so on – into hypothetical functional networks to explain what is going on.
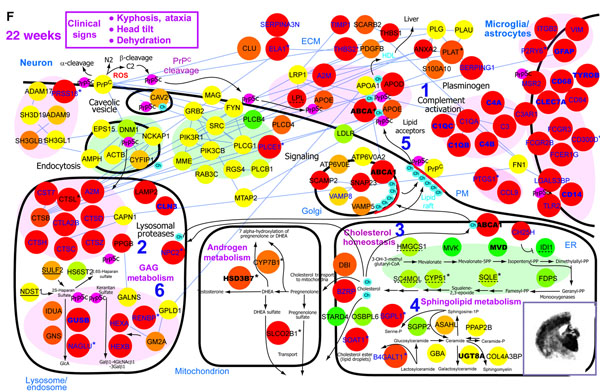
This all ends up producing giant graphs of gene names in bubbles pointing to one another, as depicted in Fig 4:
Which isn’t always the most accessible thing in the world, although Hwang’s is perhaps the most elegant I’ve seen. But even if the pictures don’t speak to you, Omenn 2009 explains why this study is a big deal: instead of getting flummoxed by the variation in prion strains, genetic factors in the host that influence susceptibility, and so forth, this study uses all that information to its advantage, finding the core of what’s always involved in prion disease biology across all those factors. Omenn also converts some of what’s in the graphs to more of a story:
Visualization…provides a dynamic scheme for the processes that characterize the molecular conversion of benign prion protein (PrPC) to disease-causing PrPSc isoforms accumulating in lipid rafts, followed by the three stages of neuropathology: synaptic degeneration, activation of microglia and astrocytes, and neuronal cell death.
But if you don’t agree with that narrative, you’re welcome to spin your own: all the data from Hwang’s study are freely open to the public in the Prion Disease Database.