Kamentsky 2011 - Improved structure, function, and compatibility for CellProfiler: modular high-throughput image analysis software
Bioinformatics. 2011 Apr 15;27(8):1179-80. Epub 2011 Feb 23.
Improved structure, function and compatibility for CellProfiler: modular high-throughput image analysis software.
Kamentsky L, Jones TR, Fraser A, Bray MA, Logan DJ, Madden KL, Ljosa V, Rueden C, Eliceiri KW, Carpenter AE.
Abstract:
There is a strong and growing need in the biology research community for accurate, automated image analysis. Here, we describe CellProfiler 2.0, which has been engineered to meet the needs of its growing user base. It is more robust and user friendly, with new algorithms and features to facilitate high-throughput work. ImageJ plugins can now be run within a CellProfiler pipeline. AVAILABILITY AND IMPLEMENTATION: CellProfiler 2.0 is free and open source, available at http://www.cellprofiler.org under the GPL v. 2 license. It is available as a packaged application for Macintosh OS X and Microsoft Windows and can be compiled for Linux. CONTACT: anne@broadinstitute.org SUPPLEMENTARY INFORMATION: Supplementary data are available at Bioinformatics online.
This two-page paper exists chiefly to explain the improvements made in CellProfiler 2.0 (released in 2011?) versus CellProfiler 1.0. In brief:
- Uses open source Python instead of proprietary Matlab.
- Object-oriented, modular
- Interoperable– bridges to C and Java / ImageJ
- Better performance, better algorithms, better UI, offers a batch mode for high throughput
- Can output analysis results directly to MySQL or SQLite
- “For neuron image analysis, CellProfiler 2.0 includes operations to enhance neurites and to measure their branching, and algorithms for neuron-specific metrics are in development.”
- “In our own research, we have used third-party ImageJ plug-ins via CellProfiler to enhance neurites in images (Supplemental Figure 1A)”
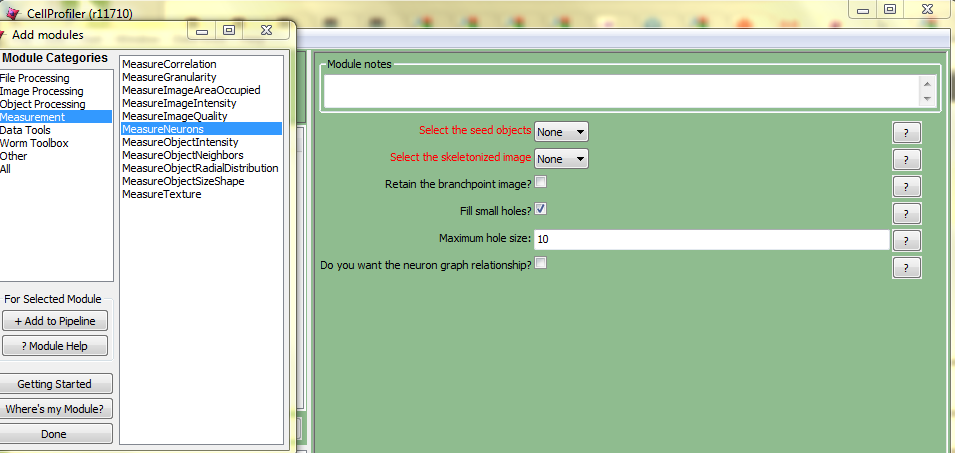
There’s no more detail about either of these. The “operations to enhance neurites and to measure their branching” in the first bullet presumably refer to the MeasureNeurons module which I see in my copy of CellProfiler:
As you can see, the UI is fairly cryptic about what measurements you’ll be getting out of this module. It will take me some serious time to figure out how to use this and how to make sense of the output.
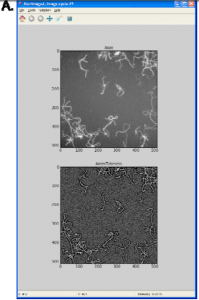
As for the second bullet, here is Supplementary Figure 1A (See also the whole supplementary data here):
The caption for this figure: “(A) CellProfiler 2.0 can run ImageJ plugins. Here, the results of running an ImageJ plug‐in, Tubeness, within a CellProfiler pipeline are shown (contrast‐enhanced, http://www.longair.net/edinburgh/imagej/tubeness/)”
From the Tubeness link: “This plugin filters a 3D image stack (or 2D image) to produce a score for how “tube-like” each point in the image is.”
So far, still nothing about recognizing astrocytes in particular, but I bet this Tubeness plugin will prove useful.