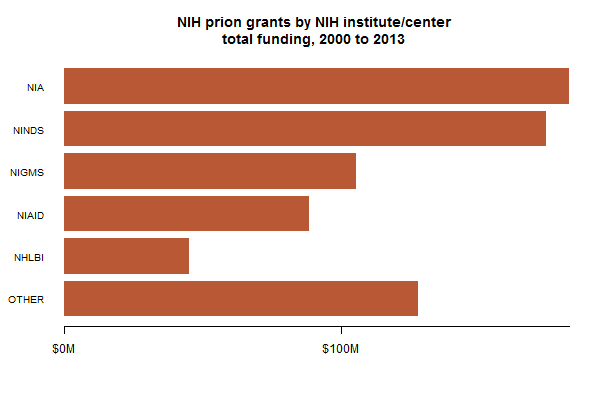Trends in NIH-funded prion grants
From 2000 to 2013, the NIH funded 2,443 grants containing the keyword “prion”, for a total value of $723 million. What can we learn about the state of public funding for prion research by mining these grants?
A few days ago I discovered the NIH RePORTER, a tool that makes certain basic information (not full text – for that you need FOIA) available to the public about every NIH grant funded in the past 25 years. This includes title, project abstract, principal investigator, institution, year, and budget among other things. A search for “prion” gave me a CSV file of 3,017 grants since 1990, and I’ve now spent a few days of quality time playing with it.
This analysis should start with several caveats. Although the dataset stretches back 25 years, not all its attributes do: project abstracts are only included from 1997 on, budgets are only included from 2000 on, and intramural grants are only listed from 2006 on. The data aren’t easy to navigate, either. There’s no proper README with exact column definitions – the closest is this About page, so I’ve been left to make some guesses as to the best interpretation of the data. For example, budgets are stated to be per year, not total for the lifetime of the grant, except for “multi-year funded” (MYF) grants, yet as far as I can tell there is no way to tell from the dataset whether a grant is MYF or not. So when I consider value of grants, should I be multiplying by the budget by the number of years, or not? Surprisingly this doesn’t make that big a difference in the analysis, because the vast majority of entries in the table are for 1 year only. That in turn is in part because some grants are duplicated across multiple rows with only the year changed – there are 3,017 rows but only 2,989 unique grant applications. This raises another issue: when analyzing the text of grants, should duplicates be removed (to avoid double-counting the same abstract) or left in (on the logic that if the grant was renewed, it deserves to be double counted)?
Where these sorts of choices matter in the analysis, I have tried to note this below. Otherwise, if you want to see exactly what I did, the code lives here. Blog comments on whether or not I took the right approach are very welcome. I want to dive into results here, so I’m not going to walk through the R code step by step like usual, other than to acknowledge that the wordclouds were created using the wordcloud R package by Ian Fellows, and that this XKCD example was helpful in learning how to use the package.
grants and funding over time
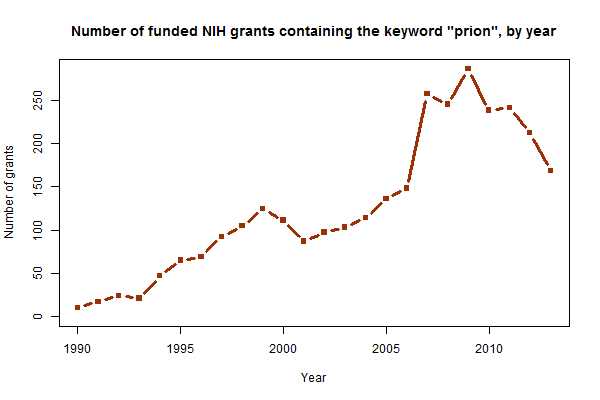
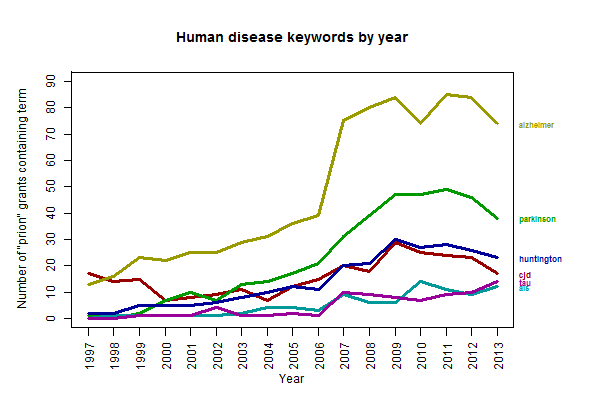
As seen above, the number of funded grants containing the word “prion” has risen epically over the past 25 years. It’s foundered a bit in the past couple years, though for what it’s worth that’s probably just the flat (in nominal terms) or declining (in real terms) NIH budget in general and not anything to do with prions in particular. The more interesting question is what’s driving the growth. Remember, the whole dataset I’m using is just based on a keyword search for “prion,” not on any more intelligent classification than that. I suspected it was probably not the case that NIH was approving more grants for PrP research than it used to, but rather probably that more applicants are calling their particular disease of interest a “prion” disease than they used to. Indeed, this graph provides some support for that:
The number of “prion” grants mentioning CJD has increased a bit but has been fairly steady. Meanwhile, the other diseases have soared – alzheimer over took CJD in 1998, parkinson did so in 2003, huntington in 2007, and tau and als look set to do so in the next year or two. To be clear, what the above plot shows is the number of funded grants mentioning both prion and the given keyword each at least once, so this doesn’t mean all of these grants are specifically focused on studying autocatalytic templating of these other proteins – some PIs may just be mentioning the prion hypothesis offhand. But still, this is quite a change. By the way: I only plotted human diseases above, but “yeast” has also been on the rise, from 0 grants for the first few years to 64 in 2009.
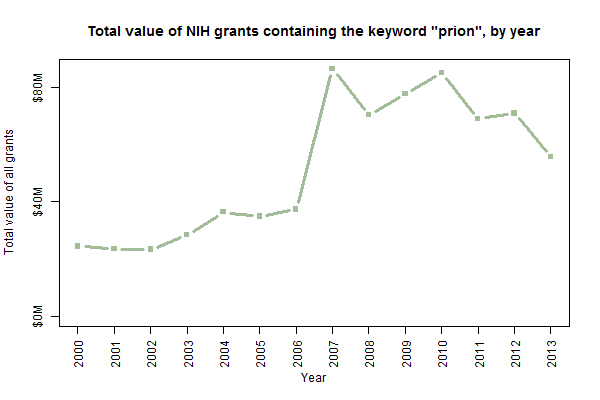
Budgets are only available in this dataset from 2000 on, but from the 14 years of data available it does appear that total spending on “prion” grants has increased in tandem with the number of grants approved:
I don’t have a good explanation for the huge uptick in 2007. That’s the first year for which intramural grants are included in this dataset, but I tried re-making this plot without the intramural grants and it didn’t look very different. I’m not sure if that was just the year everyone started saying “prion” or if it’s an artifact of the dataset.
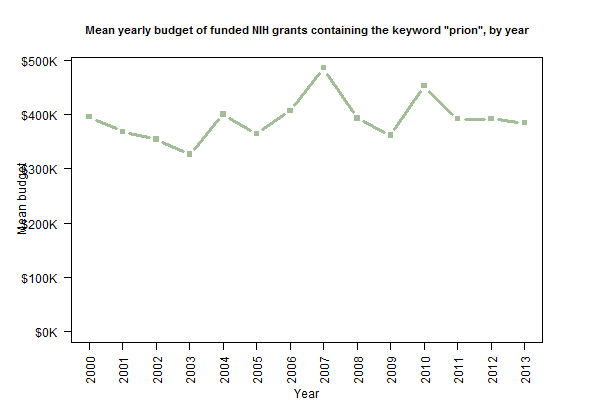
From the plots of the number of grants and the total value of grants, you can probably eyeball out the fact that the average grant value has been pretty constant. And indeed, it has been reasonably steady around $400K/yr.
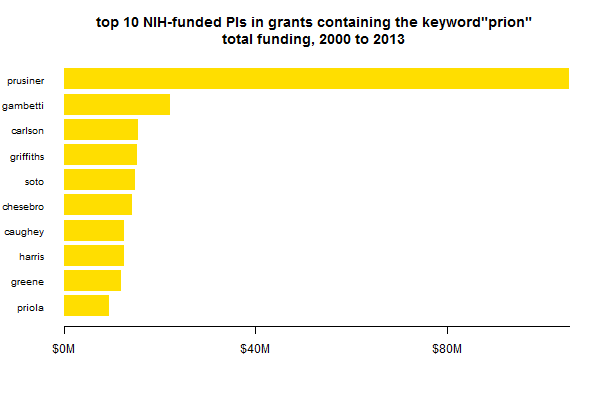
Next question: who’s been getting all this money? Here are the top 10 principal investigators in terms of total value of all grants containing the word “prion” awarded since 2000:
My first question when I saw this was, who on earth is Griffiths?? Gary Leonard Griffiths is the head of the Imaging Probe Development Center at the National Heart, Lung and Blood Institute, and is listed as PI on five nearly identical intramural grants for $3M each, supporting the IPDC over 2007-2010. Each grant names this list of IPDC-supported projects, which includes Gerald Baron‘s work on fluorescent probes for PrP trafficking. In sum: no, Griffiths is not exactly a prion PI like the other nine. His presence in the list is an unfortunate symptom of my using a simple keyword search to define the who to include.
This plot is one case where the way that you count budgets (see discussion above; was I supposed to multiply by years or not?) actually makes a big difference. Pierluigi Gambetti has an abundance of grants with multi-year durations, and if those are multiplied through, then he comes out closer to $50M over this 14-year time span.
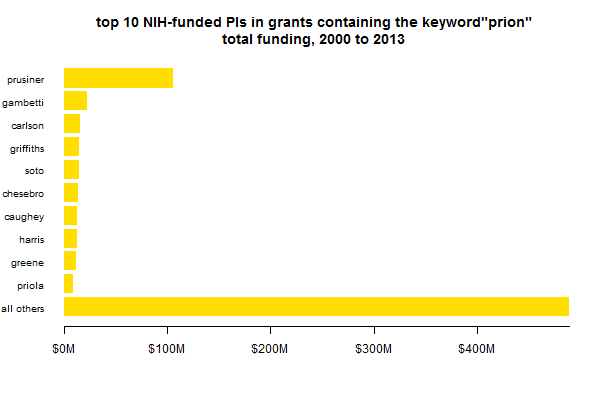
Either way, the above plot gives the appearance that Stanley Prusiner commands a really large share of prion funding. Which is partly true, but also vastly exaggerated by comparing only the top 10. There are a LOT of PIs getting prion funding (or at least mentioning the word prion once in their grants), and they add up:
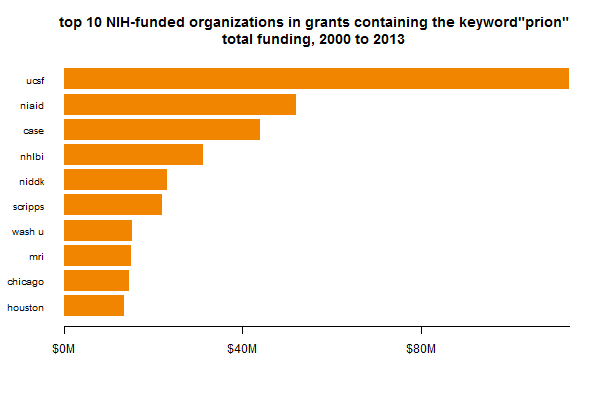
Another thing that might be slightly misleading about these plots is that it separates out PIs, not the research centers in which they work. For instance, Kurt Giles has some of his own grants within Prusiner’s center, which aren’t counted in the Prusiner bar above. What’s the total amount of funding in each center, then? If we group by center rather than PI, we get a slightly different picture:
UCSF is still #1, but NIAID (dominated by Caughey + Chesebro + Priola) appears much larger.
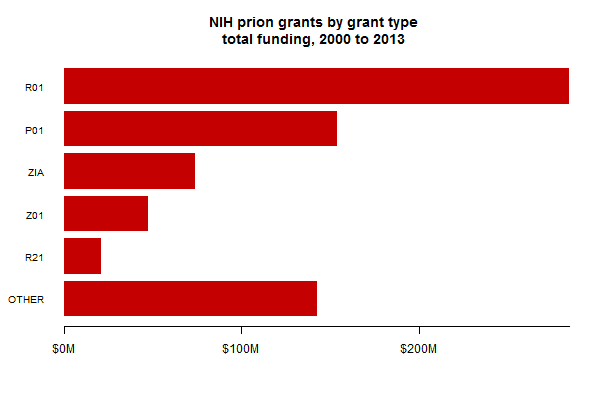
I also broke down the funding by the type of grant awarded. This looks very different from the breakdown by PI (or research center), where the top 10 put together are just a fraction of the total. About 40% of all prion funding is in the form of R01s:
Note that anything that starts with a “Z” is intramural funding.
I also had a look at which specific NIH institutes are doling out the funding:
For those who (like me) haven’t been following the NIH their whole lives, those top four stand for the National Institute on Aging, National Institute of Neurological Disorders and Stroke, National Institute of General Medical Sciences, National Institute of Allergy and Infectious Diseases. Prions, it seems, wear many hats: at times neurodegenerative disorders associated with aging, at times infectious, at times an interesting basic science question.
I’ve talked with collaborators about their grant applications a few times and it has always been desperately unclear to me how one is supposed to decide which institute to submit a prion-related grant to. This isn’t a problem in all diseases – if you’re writing a Huntington’s disease grant, for example, you usually go through NINDS, and there’s even a clear program officer to send it to (Marg Sutherland). For those of us working on (PrP) prion diseases, there is no single destination at NIH for a grant application.
In trying to get a sense of what sorts of research are funded by each of the NIH institutes, I thought perhaps I’d uncovered a perfect use case for that oft-abused visualization tool, the wordcloud. So I cooked up a “comparison cloud” for each of these top 4 institutes – NIA, NINDS, NIAID and NIGMS, where the word sizing reflects how much more often a word is used in one institute’s grants than in the other three institutes’ grants. I printed the name of each agency in the corner next to its cloud, see if you can guess which one is which before zooming in to check:
Couple of quick disclaimers. Word frequency was considered as the number of distinct grants in which the word appears at least once, but some grants were duplicated in multiple rows and in other cases the grant wasn’t exactly the same but the text was very close. Still, I just counted everything in the dataset as it was, thus doing some double counting (see discussion above). Secondly, the math here ended up a bit arbitrary. I wanted to size words by the ratio of how frequently they’re used in X institute vs. how frequently in the other 3, but you can’t divide by zero and also you want some way of “smoothing” out the results so that something that’s used 100 vs. 4 times matters more than something used 4 vs. 0 times. In the end I added 3 (totally arbitrary) to both the numerator and denominator, and normalized by the total amount of text in each agency’s grants. So you can think of the word sizing as reflecting some combination of its frequency and its degree of enrichment.
Despite the limitations, this was a moderately informative exercise. NIGMS seems to be funding a lot of the basic science, yeast and bacteria work, bioengineering: switches, biopolymers, computer simulations, eschericia, hsp100, ribosome, termination. NIAID is concerned in large part with infection, disinfection, and veterinary research (routes, horizontally, tongue, cwd-infected, sterilization, veterinarian). NIA is funding some animal modeling (transgene, huprp), translational work and drug discovery (quinacrine) and many of the “new” prion diseases (apolipoprotein, a-beta, synuclein). NINDS I didn’t get as much of a clear read from, and I’m also not sure why so few words turned up as being enriched.
I also noticed some quirks in the above wordclouds that don’t necessarily reflect anything about the NIH institute per say, but more about the top investigators who are funded by that institute. For instance, francisco in NIA is a clear giveaway: a large portion of the funding for UCSF IND (Prusiner), and all the funding for UCSF MAC (Geschwind), come through NIA. If you’ve been around the prion field a while, you’ll also notice some subtler signs that just reflect different people’s preferences regarding terminology. The top words in NIAID’s cloud are prp-res and prp-sen, operational terms which Byron Caughey favors over PrPSc and PrPC.
That observation gave me another idea, that perhaps it would be interesting to make similar comparison clouds for each of the top-funded PIs, to see which terms they mention more often than their colleagues. Here I did the math a bit differently than above: whereas for NIH institutes I counted the number of grants in which a word was used at least once, for PIs I counted the total occurrences of a word. As above, I compared each of these PIs to the other top-funded PIs, not to the whole pool of grantees.
Another disclaimer: Caughey and Chesebro’s wordclouds ended up smaller than everyone else because there just wasn’t as much text of theirs to mine, partly because intramural grants weren’t included in the dataset until 2007.
I’ll stop here at the risk of abusing this whole wordcloud concept. These images certainly aren’t analyses of any sort, just some descriptive statistics, really – but they sure are fun to look at.