Molecular Biology Workshop: PyMOL
These are my notes from the PyMOL workshop in Harvard’s BCMP 200: Molecular Biology course, delivered by Jason Key on September 15, 2014.
How to teach yourself PyMOL
Open the GUI, go to File > Log and select a path for a text file. Then do whatever you want in the GUI. The corresponding commands will be written to the file.
You can also type help <cmd>, for instance help color to get a man page for each command. To see available options, you can type <cmd> and then hit Tab, for instance color + Tab will show all available colors.
Exercise 1: secondary structure
Retrieve the structure of the T4 lysozyme and view as cartoon.
fetch 3fa0
hide everything
show cartoon
Also, in the GUI, go to Display > Sequence on to see the amino acid sequence across the top of the view window.
How many β-sheets? To count easily, color the helices red and sheets yellow:
color red, ss h
color yellow, ss s
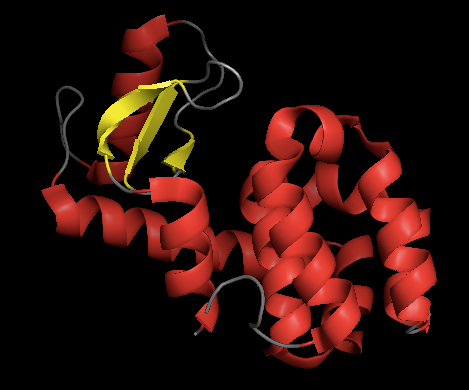
How long is the longest helix? Go to Wizard > Measurement and click on the atoms at the ends of the longest helix.
What is the angle between this helix and the other helix to its C terminal? Go to Wizard > Measurement, and in the Measurement menu that opens up in the lower right, change the measurement mode to “angle”. Then click on three representative atoms. As you keep making measurements (or trying and failing to click the right atoms), your screen will get cluttered. Click “delete all measurements” in the lower right to get rid of this clutter.
Exercise 2: sticks and lines
What is the distance across salt bridge between E22 and R137? What are the bond angles of bonds around this bridge?
hide sticks
show sticks, resi 22
show sticks, resi 137
Now with only the sticks for these two residues showing you can more easily find the nearest atoms on the two side chains and use the measurement wizard to draw a line between them. If you zoom way in on the sticks, you can also select sets of 3 atoms to measure the angles between them.
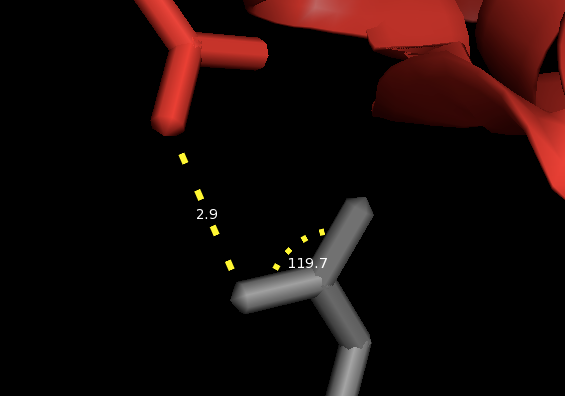
What are the overall dimensions of the T4 lysozyme? You can type zoom to see the whole thing, then just eyeball what are the farthest-apart atoms and
Exercise 3: spheres and surfaces
Which of the leucine side chains (L15, L33, L99 and L121) are buried? Which are solvent-accessible? You can tackle this by showing only these side chains under a transparent surface.
hide everything
show surface
color gray
color red, ss h
color yellow, ss s
set transparency, .5
show sticks, resi 15+33+99+121
Now by dragging it around you can more easily see which side chains are on the surface. Here, L15 (lower left) appears to be on the surface, the others look buried.
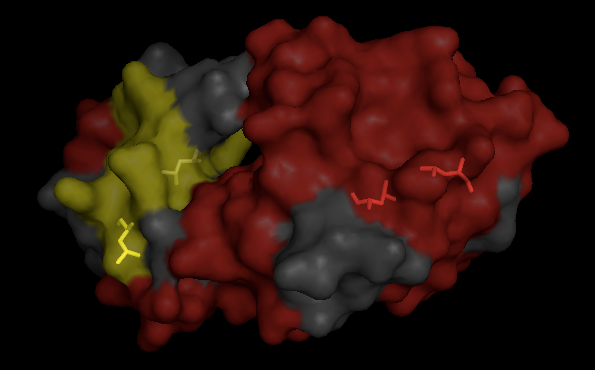
Exercise 4: solvent molecules
HOH 208 refers to a water molecule encoded as being residue #208. Which side chains coordinate this solvent molecule?
First, show this molecule:
show spheres, resi 208
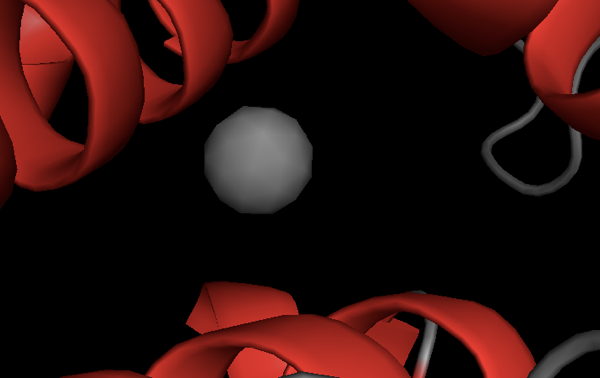

Exercise 5: ligand binding
Suppose we want to show our structure lined up in 3D space with a structure of the same enzyme bound to a ligand. Load both molecules:
fetch 148l
align 3fa0, 148l
hide everything, 148l
color blue, 148l
This lets you compare the structures.
Then in the GUI you can go to the A (actions) menu for 148l, and select presets > ligand sites > cartoon.
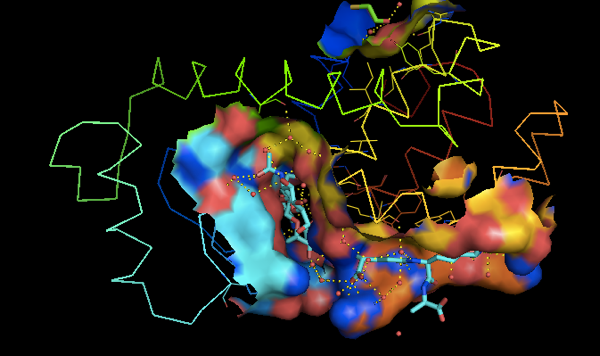
Exercise 7: crystal packing
To show adjacent molecules in order to see how the protein crystallized, go to A > generate > symmetry mates > within 4 Å
Exporting a movie
To make a movie, go to Scene > Store > F1 to store the first view, then F2 for the second one, F3 for the third, etc. Then when you hit F1, F2, F3 it will zoom elegantly between the different views. You can program this and add some motion in each step, for instance with Movie > Program > Scene loop > X-Rock > 2 seconds each. Then go to File > Save movie as… > MPEG.
