Genetics in medicine 06: Pitt-Hopkins syndrome
These are my notes from week 6 of Harvard’s Genetics 228: Genetics in Medicine from Bench to Bedside course, held on March 6, 2015. Lectures delivered by Dr. Nancy LeGendre, Dr. Ron Thibert, Dr. David Sweetser, and Dr. Steve Haggarty.
Nancy LeGendre: Introduction to Pitt Hopkins syndrome
Introduction
Dr. LeGendre is both a scientist and a parent of two daughters with Pitt-Hopkins syndrome. As she explains in this amazing blog post, finding the molecular basis of her daughters’ syndrome took 19 years and 5 geneticists at MGH. In 2012, exome sequencing finally enabled identification of a de novo Q383L variant in TCF4 in both daughters, which has been determined to be causal. This variant was presumably germline mosaic in one of the parents. Most other individuals with Pitt-Hopkins have CNV deletions in this region of chromosome 18.
Pitt-Hopkins syndrome is associated with developmental delay or regression, lack of speech, certain repetitive behaviors, abnormal breathing (either breath-holding or hyperventilation), a progressively increasing gap between chronological and developmental age, and a constellation of physical symptoms. It is sometimes described as an atypical autism, and often classified under “Pervasive developmental disorder, not otherwise specified” (PDD-NOS). Individuals with Pitt-Hopkins syndrome tend to have a very happy, positive demeanor, reminiscent of individuals with Angelman syndrome.
This year, Dr. LeGendre helped to launch the world’s first Pitt-Hopkins clinic at MGH.
Mutations in some other genes, such as NRXN1, partially phenocopy Pitt-Hopkins syndrome.
All cases of Pitt-Hopkins are caused by heterozygous mutations or deletions of TCF4. Genetic testing for Pitt-Hopkins is very new, and we still have little information on the prevalence of the disorder.
Some of the symptoms of Pitt-Hopkins are treatable. For instance chronic constipation is treated with laxatives, usually MiraLax®. Some patients exhibit improved breathing if treated with acetazolamide (Diamox™) or caffeine.
Ron Thibert: Angelman/Rett-like syndromes
Pitt-Hopkins is one of many disorders that bear phenotypic similarity to Angelman and/or Rett syndrome. Because Angelman and Rett were the first two disorders in this class to be named, the other disorders are collectively referred to as “Angelman/Rett-like syndromes” [reviewed in Tan 2014]. Symptoms of these disorders can variably include:
- Seizures
- Distinctive facial features
- Lack of speech
- Developmental delay
- Happy disposition
- Abnormal breathing
- Stereotypic hand movements
Individuals presenting with Angelman/Rett-like syndrome can now be offered gene testing panels plus array CGH to identify any of the known mutations:
| mutated gene(s) | legacy name of disorder |
|---|---|
| UBE3A | Angelman |
| MECP2 | Rett |
| TCF4 | Pitt-Hopkins |
| CDKL5 | “atypical Rett” |
| FOXG1 | “atypical Rett” |
| SLC9A6 | Christianson |
| ZEB2 | Mowat-Wilson |
| ARX | |
| MEF2C | |
| PCDH19 | |
| CNTNAP2 | |
| NRXN1 | |
| MBD5 | |
| EHMT1 | |
| 22q13.3 deletion | Phelan McDermid syndrome |
Before the genetic basis of these disorders was discovered, their diagnosis was based on lists of criteria, where a patient was said to have been diagnosed if they met, say, 2 major criteria and 1 minor criterion. This system was incredibly sloppy. Many of the genes are associated with heterogeneous clinical presentations, and these overlap between genes.
One of our classmates recommends Dr. William Gahl’s TED talk, which touches on regulatory obstacles to treating rare diseases.
David Sweetser: Clinical and molecular aspects of Pitt Hopkins syndrome
TCF4 is haploinsufficient. Causal mutations for Pitt-Hopkins syndrome include missense, splice site, nonsense, truncating frameshift, ORF-extending frameshift, balanced translocations, and CNV deletions. Although presentation varies a fair amount between individuals, all mutations appear to be equivalent, i.e. there is no obvious genotype-phenotype correlation. The TCF4 protein exists as a dimer, binds DNA, and acts as a transcription factor in the Wnt/GSK3β signaling pathway. It binds the Ephrussi-box or E-box, which has nucleotide sequence CANNTG. TCF4 is known to regulate transcription of >10 other genes, at least two of which (NRXN1 and CNTNAP2) are also implicated in Angelman/Rett-like syndromes. See also the reviews [Sepp 2012, Sweatt 2013].
Homozygous Tcf4 is perinatal lethal in mice [Korinek 1998]. TCF4 in ExAC has several supposed LoF mutations, but they are all either in the first two exons, which have low coverage and low allele number, or at the very very end of the protein.
Steve Haggarty: Therapeutic discovery
Dr. Haggarty’s lab uses patient iPS-derived neurons as a model for studying CNS disorders and developing therapeutics, an approach reviewed in [Haggarty & Perlis 2014]. Drug discovery requires a staggering number of cells. Consider that a 250,000-compound screen in duplicate in 384-well plates requires about 5 billion neurons. In order to produce neurons at this scale, iPS are first differentiated in to neural progenitor cells (NPCs) and then expanded before being terminally differentiated into desired neuronal subtypes. Dr. Haggarty’s lab has used this system to discover compounds that affect the Wnt/GSK3β signaling pathway, and to examine their effects on gene transcription [Zhao 2012].
There are a number of reasons to focus on TCF4: not only do protein function-disrupting variants cause Pitt-Hopkins syndrome, but common intronic variants are also associated with schizophrenia in GWAS. One hypothesis holds that a group of chromatin-remodeling factors, which includes neurodevelopmental disorder genes such as CHD8, are involved in regulating expression of TCF4.
Journal club: Crebinostat: a novel cognitive enhancer that inhibits histone deacetylase activity and modulates chromatin-mediated neuroplasticity [Fass 2013]
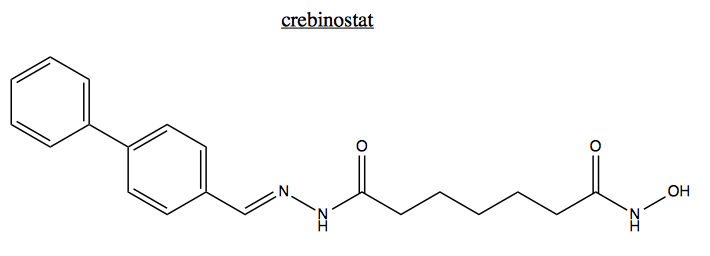
For journal club this week we discussed a paper from Steve Haggarty’s group introducing crebinostat as a histone deacetylase inhibitor [Fass 2013].
