Exome sequencing pipeline using GATK
overview: This post documents a pipeline for human exome sequencing using GATK. The prefatory remarks from the bowtie2/samtools exome pipeline I’ve posted apply to this pipeline as well:
This pipeline is for analyzing human exome data. It assumes you have paired-end reads in the form of FASTQ files from your sequencing company and that your goal is to align these to the genome and then call variants for further analysis. Below I will include code and estimated running times for every step. For the record, I’ve got 50 samples with paired-end reads, so 100 FASTQ files, ranging from about 15 to 30 million reads each, and I am running this on an HP Red Hat Linux cluster with 8GB memory and 8 cores per node. But IMHO, the details aren’t important– your running times will obviously be different than mine, but this will give you a sense of the order of magnitude of how long these things will take. That’s helpful because then you know whether to kill them if they’ve been running too long, and you can set running time limits on your jobs (using bsub‘s -W option if you’re on an LSF scheduler) which will help keep your priority nice and high on shared resources.
motivation: I recently developed an exome sequencing pipeline using bowtie2 and samtools. But because GATK is such an industry standard tool and seems to have a stream of constant improvements to offer, I have also been interested in getting an exome pipeline up and running using GATK. As I mentioned last week, GATK isn’t quite an end-to-end solution as it still relies on BWA and Picard for alignment and pre-processing, and it is all too easy to run into compatibility problems when calling BWA, Picard and GATK separately. For this reason, I was attracted to Queue, a framework that allows you to create pipelines in Scala that stitch together various tasks. I don’t know Scala, and was not super keen on learning a new programming language just to use GATK, but Broad provides some Queue pipelines, including a DataProcessingPipeline, that you can (almost) use out of the box. Making use of the DataProcessingPipeline as a starting point I was finally able to run GATK on my data. update 2013-06-04: Broad has since deprecated the DataProcessingPipeline and pulled it from their site; below I provide the version of it that I had working successfully in this pipeline. disclaimer: I didn’t want to learn Scala to do this, but doing so, and writing a pipeline for Queue, would clearly be the right way to pipeline GATK. Instead I used a combination of calls to Queue using existing scripts along with direct calls to GATK and submitted all these things as separate jobs to my LSF scheduler. This worked for me, but I’m told you can get much better performance by letting Queue manage LSF jobs for you so it can parallelize stuff within GATK. If I end up doing a lot more exome sequencing I guess I’ll have to look into it.
0. prep steps Before you can do anything, make sure you’ve got GATK, Queue, DataProcessingPipeline and the hg19 resource bundle. Download GATK and Queue here (you need to register for an account first) and find the text of the latest DataProcessingPipeline.scala here. update: as of 2013-06-04: Broad has deprecated the DataProcessingPipeline. Because my pipeline depends on it, I am providing the version of DataProcessingPipeline.scala that worked with my pipeline: DataProcessingPipeline.scala I cannot guarantee compatibility with newer builds of GATK and Queue. after you download, rename to remove the .txt extension. To get the resource bundle, visit Broad’s GSA public FTP server; as of September 12, 2012, the latest version is 1.5 (I’m using hg19) and you can get it, checksum it (thanks @a) and decompress it via the unix command line as follows:
wget -r ftp://gsapubftp-anonymous@ftp.broadinstitute.org/bundle/1.5/hg19/ # download all files in this directory ls *.md5 | awk '{print "sed -i \x27s|/humgen/gsa-scr1/pub/bundle/1.5/hg19/||\x27 " $0}' | bash # replace the absolute paths in all the md5 files ls *.md5 | awk '{print "md5sum -c " $0}' | bash # check all the md5sums - if any are not 'OK' then stop here. ls *.gz | awk '{print "gunzip " $0}' | bash # decompress all the gz files
A note about the above. Broad uses absolute paths in its md5 files so md5sum -c will fail if you try to use them as is. In the second line above, I use sed to remove the absolute path information from each file. update 2013-02-05: I have heard you may get faster alignment results in step 4 if you use the regular human reference genome plus the decoy genome as your reference. Haven’t tested it myself to see how much faster. end updateThe resource bundle does not include the bwa indices for hg19, so you’ll have to create those yourself:
bwa index ucsc.hg19.fasta
Which takes about 3 hours. Also a general note: before you go ahead and actually run the following steps on your full FASTQs with millions of reads, you might want to try out the pipeline on much shorter files. I used head -40000 1_1.fq > subset/1_1.fq to create shorter FASTQs with just 10,000 reads each to use while debugging my pipeline. The running times I give below are for my full-size files, not for the shortened test version. For steps 5 and 7 you will also want a BED file that lists the regions you targeted when you did your exome capture. If you used the Illumina TruSeq exome capture kit, the official BED file for it is here but you need to log in to download; if you trust things people post on the internet, someone has uploaded a free copy here. If your exome capture kit was NimbleGen SeqCap, the relevant page is here and the actual files are here: v2.0 and v3.0. (Note that the NimbleGen v2 BED files contain two tracks (see under “File Descriptions”); you’ll want to delete all the rows from the first track to avoid redundancy, which you can just do in a text editor. Search for “tiled_regions” and you’ll find the line dividing the two tracks.)
1. check fastq validity: checksum or gunzip To get your raw data, you probably downloaded .fq.gz files from your sequencing company. These files are huge, so there’s plenty of opportunity for them to get corrupted during the download. In case this should occur, you want to check for errors before starting your pipeline so you don’t waste CPU cycles processing the data before hitting a corrupted line and needing to backtrack. Ideally the company will give you a checksum file and you can use md5sum or sha1sumto check for corruption. In my case, this was not provided, so I found it easiest to just gunzip the files before starting– gunzip will catch any invalid compression blocks.
gunzip -c 1_1.fq.gz 1_1.fq
3 minutes per .fq.gz file If there’s a corrupted file you’ll get a message like invalid compressed data--format violated. Important note: gunzip’s default behavior is to remove the compressed file when it’s done; to be safe you should probably keep it using the -c flag as I’ve done above.
2. raw reads quality control: fastqc
fastqc 1_1.fq -f fastq -o fastqc/
8 minutes per fq file I’ve provided a script to load the FastQC output data into PostgreSQL to analyze in bulk here.
3. fastq to unaligned bam: picard FastqToSam I’ve had format compatibility problemsbetween BWA, Picard and GATK before, so I want to let GATK do the alignment so I know it’s done exactly how GATK wants it done. So in step 4 I am going to have the DataProcessingPipeline be in charge of callling BWA to “redo” the alignment, so for now I just need to convert my FASTQs to BAMs without aligning. I also need to specify the appropriate @RG flags in the header, which is necessary for other tools down the line. This can all be done in one call to picard:
java -Xmx2G -jar ~/bin/picard-tools-1.74/FastqToSam.jar FASTQ=fq/1_1.fq FASTQ2=fq/1_2.fq QUALITY_FORMAT=Illumina OUTPUT=1.bam READ_GROUP_NAME=rgname SAMPLE_NAME=1 SORT_ORDER=unsorted PLATFORM=ILLUMINA PLATFORM_UNIT=pu1 LIBRARY_NAME=lbname
This took 8 – 13 minutes per pair of FASTQs, varying pretty much proportional to the number of reads.
4. alignment and pre-processing: Queue / DataProcessingPipeline Broad has provided the DataProcessingPipeline to accomplish basically all the steps of Phase I of its best practices v4. Call Java to call Queue to run DataProcessingPipeline.scala:
java \
-Xmx4g \
-Djava.io.tmpdir=/tmp/mytmpdir/ \
-jar ~/bin/gatk-2.1-5/Queue-2.1-8/Queue.jar \
-S ~/bin/gatk-2.1-5/Queue-2.1-8/DataProcessingPipeline.scala \
-bwa ~/bin/bwa-0.6.2/bwa \
-i 1.bam \
-R gatk-bundle-1.5-hg19/ucsc.hg19.fasta \
-D gatk-bundle-1.5-hg19/dbsnp_135.hg19.vcf \
-p project_name \
-outputDir processedbams/ \
-bwape \
-run
Running time ranged from 20 hours to 33 hours, varying pretty much proportionally with the number of reads. Description of parameters is here. By specifying -bwape I tell it to completely redo the alignment using bwa sampe. When DataProcessingPipeline is done, you will find a file that looks like project_name.sample_name.clean.dedup.recal.bam.out in your processedbams directory. Provided that everything worked, this is all you need going forward.
5. calculate coverage: bedtools You can do this now, or you can do it later. Now that you’ve done alignment, at some point you will want to know at what depth you sequenced your samples. bedtools coverageBed can do this, though it produces a lot more data than I need, so I pipe its output directly to tailto get a much smaller file containing the summary rows that I care about:
coverageBed -abam project_name.1.clean.dedup.recal.bam -b SeqCap_EZ_Exome_v2.bed -hist | tail -10000 - > 1.coverage
5 minutes per bam Then I wrote a Python script to read the summary rows from all my samples into a PostgreSQL table:
import sys
import csv
import psycopg2
ctsql="""
drop table if exists exome_coverage_histo;
create table exome_coverage_histo (
sample_id integer,
depth integer,
bases integer
);
"""
conn = psycopg2.connect(database="mydb", user="postgres", password="password")
conn.autocommit = True
cur = conn.cursor()
cur.execute(ctsql)
localdir = "c:/your/path/here/"
for i in range(0,50):
f = open(localdir+str(i)+".coverage","r")
c = csv.reader(f, delimiter="\t")
for row in c:
if (row[0] <> "all"):
continue
cur.execute("insert into exome_coverage_histo(sample_id,depth,bases)values(%s,%s,%s);",(i,int(row[1]),int(row[2])))
f.close()
cur.close()
conn.close()
That took about 2 minutes to run. And then you can get your summary stats with a few SQL queries:
-- Average coverage of target by sample
select sample_id, sum(depth*bases)::numeric / sum(bases)::numeric as avg_coverage
from exome_coverage_histo
group by sample_id
order by avg_coverage
;
-- Average coverage of target across all samples
select sum(depth*bases)::numeric / sum(bases)::numeric as avg_coverage
from exome_coverage_histo
;
-- Percent of target covered at 8x or greater, by sample
select sample_id, sum(case when depth >= 8 then bases else 0 end)::numeric / sum(bases)::numeric as fraction8x
from exome_coverage_histo
group by sample_id
order by fraction8x
;
-- Percent of target covered at 8x or greater, across all samples
select sum(case when depth >= 8 then bases else 0 end)::numeric / sum(bases)::numeric as fraction8x
from exome_coverage_histo
;
In step 7, I will create modified BED files which include a 10bp buffer on either side of targeted regions, for the purpose of calling variants. As far as I can tell, though, most articles in the literature refer to “coverage of target”, not “coverage of target plus buffer”, which is why I used the original BED file for this step.
6. reduce reads: GATK ReduceReads This is a clever tool previously mentioned herewhich can theoretically reduce BAM size (and therefore processing time for downstream steps) by 10 – 100x. It’s not bundled into DataProcessingPipeline and I found it easier to just call it separately than to learn enough Scala to add it in.
java \
-Xmx8g \
-Djava.io.tmpdir=/tmp/mytmpdir/ \
-jar ~/bin/gatk-2.1-5/GenomeAnalysisTK.jar \
-T ReduceReads \
-I project_name.1.clean.dedup.recal.bam \
-R gatk-bundle-1.5-hg19/ucsc.hg19.fasta \
-o project_name.1.clean.dedup.recal.reduced.bam
Running time for most samples ranged from 2 to 4 hours, but variance was high. Several samples were in the 5 hour range, and two outliers clocked in at 8 and 15 hours. The BAMs were each reduced to about 40% of original size (this is for exome sequencing with coverage of target in the 30x-50x range). The input BAMs were in the 7 – 10GB range and the output BAMs were in the 2.5 – 4GB range. ReduceReads currently suffers from memory allocation issues as discussed in the Broad GATK forums, though this may be fixed soon. It crashed with 4GB memory but when I ran it with 8GB memory (-Xmx8g argument to Java above) it successfully completed on all my files. If it crashes you may need to start this step over again for those samples with even more memory. Important note #1: if you are running your pipeline on shortened FASTQs for debugging purposes (see step 0 above), you’ll find that this step not only fails to reduce BAM size, it may even increase it slightly. Not to worry. When you shortened your FASTQs you caused there to be something like .1x coverage of the exome instead of 100x or whatever you had to start with. ReduceReads can only compress where reads overlap, which is almost nowhere when you have just .1x coverage. When you come back later and run this pipeline on your full dataset, this step will reduce the file size. Important note #2: bedtools doesn’t know how to read the special annotation that Broad uses to note synthetic reads in reduced read BAMs. In regions where one read in the reduced BAM is standing in for tens of reads in the original bam, the bedtools coverageBam tool will interpret it as just one read. Therefore to accurately calculate coverage you need to use the original BAMs from the end of Step 4, i.e. project_name.1.clean.dedup.recal.bam. If you decide to do step 5 later rather than earlier, just be sure to use the non-reduced BAMs as input.
7. call variants: GATK UnifiedGenotyper Now you’re ready to pile up the reduced read BAMs from all your samples and call variants on them. There are a variety of ways to parallelize this, including using Queue– see this conceptually helpful forum discussion. If you don’t parallelize, see this graph of how long UnifiedGenotyper will take in serial: it’s a long time (though I think this is for whole genome, not exome). UnifiedGenotyper’s operations are actually independent by site (not just by chromosome), which means you can break your analysis into (almost) arbitrarily small pieces in order to parallelize it. To call variants on only a subregion of the genome, you can call UnifiedGenotyper with the -L flag, for instance, -L chr1. But wait, we’re doing exome sequencing here, so we also want to restrict our analysis to the exome, using -L targetedregions.bed. While some reads will align to non-targeted regions, coverage there will be so low you will not be able to reliably call many variants anyway, so you may as well save time and not bother genotyping those regions. But, to add another wrench to the issue, the BED files provided by companies that make exome capture kits seem to specify only the exons themselves, whereas coverage on the immediately adjacent splice sites will very likely be good enough to call variants, so you do want to include a small buffer around your exons (SeqAnswers recommends a 10bp buffer). update 2012-10-02: Because mitochondria are so numerous in a cell, you tend to get high (off-target) coverage of mtDNA even if your exome capture kit doesn’t target any mitochondrial genes (and most don’t). Picardi & Pesole 2012 [supplement] explain more. For this reason it may be worth including the entire mitochondrial chromosome in your BED file as well. You can do this by simply adding a single line to the end of your BED file:
chrM 0 16571
But you may want to calculate coverage for the mitochondrial chromosome separately just to make sure you have enough. end update update 2012-10-17: The NimbleGen BED files are encoded for hg19. If you need to make them compatible with GRCh37, you need to do five things: remove the letters ‘chr’ from each chromosome name, remove the random contigs, remove the ‘un’ contigs, change ‘M’ to ‘MT’ for mitochondria, and change the mitochondrial base range. Here’s a bash one-liner for the first four:
sed 's/^...//' hg19_targets.bed | grep -v random | grep -v Un | sed -e 's/M/MT/' > GRCh37_targets.bed
For the last one, just open your bed file and edit the last line. hg19 recognizes bases 0 – 16571 of mtDNA, while according to GRCh37 it only goes up to 16569. If you don’t change your last line to:
MT 0 16569
then you’ll get a “array index out of range” error when you run GATK’s UnifiedGenotyper. end updateTaking all these issues into account, I downloaded an exome BED file (see step 0 above) and wrote a quick Python script to accomplish two things: (1) expand all the intervals by 10 bp on either side, clipping in cases where this would result in overlap between consecutive intervals, and (2) break the BED file into 1000-interval chunks so that I could parallelize. Here’s the script:
import sys
import csv
extrabases = 10 # size of extra window to create around targeted exons
outputbedsize = 1000 # number of intervals per output BED file to create
inbed = open("c:/your/path/exome.bed","r")
c = csv.reader(inbed,delimiter="\t")
overlapsprevented = 0
counter = 0
oldend = 0
oldchro = 'chr1'
outbed = open("c:/your/path/"+str(counter%outputbedsize)+".bed","wb")
for row in c:
chro = row[0]
start = int(row[1]) - extrabases # start 10bp before target
end = int(row[2]) + extrabases # end 10bp after target
if(chro == oldchro and start < oldend): # avoid overlap
start = oldend+1 #... by starting after the previous interval ended
overlapsprevented +=1
if(start > end): # and if one interval is entirely contained in another, just toss it
continue
outbed.write("\t".join((chro,str(start),str(end)))+"\n") # write to output bed file
counter += 1
# keep track of this iteration's values to compare for overlap on next iteration
oldend = end
oldchro = chro
if(counter%outputbedsize == 0): # split into new file every 1000 lines
outbed.close()
outbed = open("c:/your/path/"+str(counter/outputbedsize)+".bed","wb")
outbed.close()
inbed.close()
print "number of overlaps prevented: ", overlapsprevented
update 2013-02-05: I later learned a quicker way to do this without Python scripting. bedtools slopBed can add margins (ex. 10bp) on each side of your ranges, and the unix command split can separate it into separate files of 1000 lines each. slopBed uses a .genome file to make sure it doesn’t extend past the end of the chromosome, so you’ll need the hg19.genome file (or equivalent) from the bedtools github. The overall command will look like so:
slopBed -i exome.bed -g hg19.genome -b 10 | split -d -a 3 -l 1000 - part
end updateThis gave me hundreds of BED files with 1000 lines each. Then I could parallelize by running multiple jobs, each of which piles up all of my reduced BAMs from step 5 but uses only one of my BED files and thus only has a small region of the genome to be concerned about. Here’s an example call:
java \ -Xmx4g \ -Djava.io.tmpdir=/tmp/evm-gatk/ \ -jar ~/bin/gatk-2.1-5/GenomeAnalysisTK.jar \ -T UnifiedGenotyper \ -R gatk-bundle-1.5-hg19/ucsc.hg19.fasta \ -I projectname.1.clean.dedup.recal.reduced.bam -I projectname.2.clean.dedup.recal.reduced.bam [... etc. list remaining bams] \ -o 0.vcf \ --dbsnp gatk-bundle-1.5-hg19/dbsnp_135.hg19.vcf \ -glm BOTH \ -L 0.bed \ -stand_call_conf 30.0 \ -stand_emit_conf 30.0 \ -dcov 200
Running time: 2.5 to 5 minutes per 1000-line BED file, or about 16 hours total. The arguments above were chosen based largely on Broad’s best practices v4 doc. After you’ve run the above commands, you’ve got hundreds of little VCF files. Now recombine those using CombineVariants:
java \ -Xmx8g \ -jar ~/bin/gatk-2.1-5/GenomeAnalysisTK.jar \ -R gatk-bundle-1.5-hg19/ucsc.hg19.fasta \ -T CombineVariants \ --variant 0.vcf \ --variant 1.vcf \ --variant 2.vcf \ -o variants.vcf \ -genotypeMergeOptions UNIQUIFY
Which only takes about 3 minutes.
8. recalibrate variants: GATK VariantRecalibrator and ApplyRecalibration As discussed here, Broad has undergone a major effort to recalibrate quality scores so that they reflect the actual log frequency with which they are incorrect, which is what PHRED scores are supposed to mean. This functionality is split over two tools: VariantRecalibrator creates a model, and ApplyRecalibration uses the model to adjust the scores. It’s discussed in a good amount of detail on the best practices v4 doc as well as this VQSR discussion. I just used what’s recommended there, and everything was mercifully quick:
# create model for SNPs
java -Xmx8g -jar ~/bin/gatk-2.1-5/GenomeAnalysisTK.jar \
-T VariantRecalibrator \
-R gatk-bundle-1.5-hg19/ucsc.hg19.fasta \
-input variants.vcf \
--resource:hapmap,known=false,training=true,truth=true,prior=15.0 gatk-bundle-1.5-hg19/hapmap_3.3.hg19.sites.vcf \
--resource:omni,known=false,training=true,truth=false,prior=12.0 gatk-bundle-1.5-hg19/1000G_omni2.5.hg19.sites.vcf \
--resource:dbsnp,known=true,training=false,truth=false,prior=6.0 gatk-bundle-1.5-hg19/dbsnp_135.hg19.vcf \
-an QD -an HaplotypeScore -an MQRankSum -an ReadPosRankSum -an FS -an MQ -an InbreedingCoeff \
-mode SNP \
-recalFile variants.snps.recal \
-tranchesFile variants.snps.tranches
Running time: 9 minutes
# create model for INDELs
java -Xmx8g -jar ~/bin/gatk-2.1-5/GenomeAnalysisTK.jar \
-T VariantRecalibrator \
-R gatk-bundle-1.5-hg19/ucsc.hg19.fasta \
-input variants.vcf \
--maxGaussians 4 -std 10.0 -percentBad 0.12 \
--resource:mills,known=true,training=true,truth=true,prior=12.0 gatk-bundle-1.5-hg19/Mills_and_1000G_gold_standard.indels.hg19.sites.vcf \
-an QD -an FS -an HaplotypeScore -an ReadPosRankSum -an InbreedingCoeff \
-mode INDEL \
-recalFile variants.indels.recal \
-tranchesFile variants.indels.tranches
Running time: 20 seconds
# apply SNP model
java -Xmx8g -jar ~/bin/gatk-2.1-5/GenomeAnalysisTK.jar \
-T ApplyRecalibration \
-R gatk-bundle-1.5-hg19/ucsc.hg19.fasta \
-input variants.vcf \
--ts_filter_level 99.0 \
-tranchesFile variants.snps.tranches \
-recalFile variants.snps.recal \
-mode SNP \
-o variants.intermediate.vcf
Running time: 13 seconds
# apply INDEL model
java -Xmx8g -jar ~/bin/gatk-2.1-5/GenomeAnalysisTK.jar \
-T ApplyRecalibration \
-R gatk-bundle-1.5-hg19/ucsc.hg19.fasta \
-input variants.intermediate.vcf \
-ts_filter_level 99.0 \
-tranchesFile variants.indels.tranches \
-recalFile variants.indels.recal \
-mode INDEL \
-o variants.analysisready.vcf
Running time: 12 seconds
9. annotation: snpEff UnifiedGenotyper itself can provide a variety of annotations (full list at left navbar here), but these are mostly annotating information about your own data. To get information about gene names, coding regions, effects of variants and so on, you still have to use a separate tool, snpEff, as explained here. As for downloading the hg19 reference info for snpEff, the link on the snpEff download page didn’t work for me but I was able to get what I needed from sourceforge. snpEff is super easy to use; the only slightly weird thing that tripped me up is you have to run it from the directory where it is installed, instead of from the directory where your data is. Other than that it’s a breeze:
cd ~/bin/snpEff_2_0_5/ java -Xmx8G -jar snpEff.jar eff -v -onlyCoding true -i vcf -o vcf hg19 working/dir/here/variants.analysisready.vcf > working/dir/here/variants.analysisready.snpeff.vcf
Which runs in about 3 seconds. Which is pretty incredible considering it not only performs the annotation and outputs a several hundred megabyte VCF file to disk, but also creates two other handy files in the directory where you ran it. snpEff_summary.html contains an dazzling array of descriptive statistics (an absolute marvel for 3 seconds of runtime, really) and snpEff_genes.txt is a database-ready list of how various genes are affected by variants in your samples. One shortcoming of snpEff is that, as its name suggests, it exists to annotate the effect of variants. It doesn’t provide any of the non-effect-related metadata you might be interested in: SIFT scores, phyloP conservation scores, 1000 genomes minor allele frequency, and so on. For those things you’ll want to use annovar; see my previous exome pipeline (step 9) for some tips.
10. conversion to database format: GATK, python and PostgreSQL I found that the VCF file produced by snpEff was not perfectly well-formed. In particular: for rows with exactly one effect, snpEff adds an extra tab between the INFO and FORMAT fields, and for rows with more than one effect, snpEff eliminates the FORMAT field. This may be a snpEff bug, or (though I can’t imagine exactly why) this could be because I used hg19, which snpEff discourages (they prefer GRCh37, though they do still offer hg19 as an option). In any case, this makes it tough to read the data into a table. However, GATK’s VariantAnnotator appears to be capable of tolerating this issue, presumably since it doesn’t read past the INFO field of each line in the snpEff VCF file. You can use VariantAnnotator to copy the snpEff annotations from the snpEff output VCF file back into your original VCF file:
java -Xmx8g -jar ~/bin/gatk-2.1-5/GenomeAnalysisTK.jar \
-R gatk-bundle-1.5-hg19/ucsc.hg19.fasta \
-T VariantAnnotator \
-E resource.EFF \
--variant variants.analysisready.vcf \
--resource variants.analysisready.snpeff.vcf \
-o variants.analysisready.snpeff.backin.vcf
Running time: 40 minutes. Note that the -E flag just lets you specify what the name of the column containing the snpEff effects will be. You don’t need an input file named resource.EFF. Using the -E flag will give you one column with all the snpEff information in it, which means you’ve got a many-effects-to-one-variant relationship on your hands, which I relationalize into my database in just a moment. But if you’d rather not deal with the many:one complexity, you can just use the -A flag instead, which will give you only the highest-impact effect for each variant. See the relevant GATK forum discussion. Next, you can use GATK’s VariantsToTable to create something a bit more database-friendly: it can pull all the INFO and FORMAT fields out and make them each their own column. It’s still not relational, but at least it’s a proper table and you don’t need to quite as much text parsing as what I was doing before to deal with the raw VCFs from samtools. Here’s a sample call:
java -Xmx8g -jar ~/bin/gatk-2.1-5/GenomeAnalysisTK.jar \
-R gatk-bundle-1.5-hg19/ucsc.hg19.fasta \
-T VariantsToTable \
-o variants.table \
--allowMissingData \
--showFiltered \
-V variants.analysisready.snpeff.backin.vcf \
-F CHROM -F POS -F ID -F REF -F ALT -F QUAL -F FILTER -F AC -F AF -F AN -F BaseQRankSum -F DB -F DP -F DS -F Dels -F END -F FS -F HaplotypeScore -F InbreedingCoeff -F MLEAC -F MLEAF -F MQ -F MQ0 -F MQRankSum -F QD -F RPA -F RU -F ReadPosRankSum -F SB -F STR -F VQSLOD -F culprit -F set \
-F resource.EFF \
-GF GT -GF AD -GF DP -GF GQ -GF PL
Running time: 30 seconds. If your hideously long list of -F and -GF arguments isn’t identical to mine, you can retrieve it by copying the ##INFO lines out of your VCF into a text editor and then doing a bunch of replacement operations on them. Next I want to step through the table line by line, recreating a relational model as I go. I want something a bit simpler and easier to work with than the full Ensembl model, but I still want to have the data in a relational format so that it’s easy to summarize data across different attributes, drill down to individual samples, pull data out to work with in R, join to other datasets, etc. Here’s the relational model I’m aiming at (this assumes you’ve got a samples table somewhere with metadata on each of your samples). 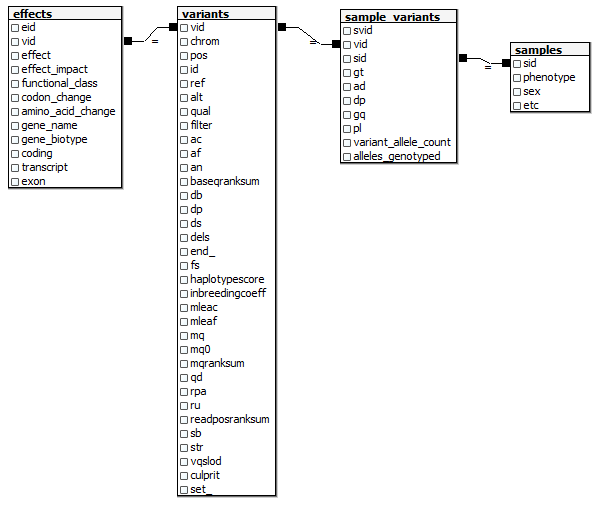 And here’s the Python code I wrote to create this model from the
And here’s the Python code I wrote to create this model from the variants.tablefile from GATK:
import sys
import psycopg2 # http://www.initd.org/psycopg/
# "set" and "end" are sql keywords, so add an underscore to them in your input file header to match this create table statement.
ctsql = """
drop table if exists vartest.variants;
create table vartest.variants(
vid serial primary key,
chrom varchar,
pos integer,
id varchar,
ref varchar,
alt varchar,
qual varchar,
filter varchar,
ac varchar,
af varchar,
an integer,
baseqranksum numeric,
db varchar,
dp integer,
ds varchar,
dels varchar,
end_ integer,
fs numeric,
haplotypescore numeric,
inbreedingcoeff numeric,
mleac varchar,
mleaf varchar,
mq numeric,
mq0 integer,
mqranksum numeric,
qd numeric,
rpa varchar,
ru varchar,
readposranksum numeric,
sb numeric,
str varchar,
vqslod numeric,
culprit varchar,
set_ varchar
);
drop table if exists vartest.effects;
create table vartest.effects(
eid serial primary key,
vid integer,
effect varchar,
effect_impact varchar,
functional_class varchar,
codon_change varchar,
amino_acid_change varchar,
gene_name varchar,
gene_biotype varchar,
coding varchar,
transcript varchar,
exon varchar
);
drop table if exists vartest.sample_variants;
create table vartest.sample_variants(
svid serial primary key,
vid integer,
sid integer,
gt varchar,
ad varchar,
dp varchar,
gq varchar,
pl varchar,
variant_allele_count integer,
alleles_genotyped integer
);
"""
effectcolnamelist = "effect , effect_impact , functional_class , codon_change , amino_acid_change , gene_name , gene_biotype , coding , transcript , exon"
neffectcols = len(effectcolnamelist.split(","))
def insertEffects(c,vid,multieffectstring):
if(multieffectstring == "NA"):
return
# strip brackets
if(multieffectstring[0] == "[" and multieffectstring[-1] == "]"):
multieffectstring = multieffectstring[1:-1]
# break up multiple effects
effectstringlist = multieffectstring.split(",")
# insert a row for each effect
for effectstring in effectstringlist:
# remove any whitespace
effectstring = effectstring.strip()
# retrive the effect from before the parens
effect = effectstring.split("(")[0]
# break the rest of the data up by pipes
restofdata = effectstring[effectstring.find("(")+1:effectstring.find(")")].split("|")
data = tuple([vid] + [effect] + restofdata)
sql = "insert into vartest.effects(vid," + effectcolnamelist + ") values(" + "%s,"*(neffectcols+1)
sql = sql[:-1]+ ");"
c.execute(sql,data)
# open database connection
conn = psycopg2.connect(database="mydb", user="postgres", password="password")
conn.autocommit = True
c = conn.cursor()
# drop and create tables
c.execute(ctsql)
# prep to read input table
inpath = "c:/your/path/here/variants.table"
infile = open(inpath,"r")
delim = "\t"
nvcols = 33
effcol = 33
samplecolstart = 34
colspersample = 5
samples = 50
# retrieve header
header = infile.readline().lower().split(delim)
variants_col_list = header[:nvcols]
linecounter = 1
vid = 0
# iterate through input file, writing to database
for line in infile.readlines():
datalist = line.split(delim)
# variants table
sql = "insert into vartest.variants("+",".join(variants_col_list)+")"
sql += "values (" + "%s,"*len(variants_col_list)
sql = sql[:-1] + ") returning vid;"
vardata = line.split(delim)[:nvcols]
for i in range(len(vardata)):
if(vardata[i]=="NA"):
vardata[i] = None
data = tuple(vardata)
c.execute(sql,data)
vid = c.fetchone()[0]
# effects table
insertEffects(c,vid,line.split(delim)[effcol])
# sample variants table
samplecols = line.split(delim)[samplecolstart:]
assert len(samplecols) == colspersample*samples, "wrong number of samples at line " + str(linecounter) + "\n" + line
for i in range(samples):
svid = 0
sid = header[samplecolstart+i*colspersample].split(".")[0]
samplecolnamelist = []
sampledatalist = []
for j in range(colspersample):
masterindex = samplecolstart + i*colspersample + j
samplecolnamelist.append(header[masterindex].lower().split(".")[1])
sampledatalist.append(datalist[masterindex])
# deal with genotype and allele count
variant_allele_count = 0
alleles_genotyped = 0
alleles = sampledatalist[0].split("/")
ref = datalist[3]
alt = datalist[4]
for allele in alleles:
if(allele == "."):
variant_allele_count = None
break
elif(allele == alt):
variant_allele_count += 1
alleles_genotyped += 1
elif(allele == ref):
variant_allele_count += 0
alleles_genotyped += 1
sql = "insert into vartest.sample_variants(vid,sid,"+",".join(samplecolnamelist)+",variant_allele_count,alleles_genotyped) "
sql += "values (" + "%s,"*(colspersample+4)
sql = sql[:-1]+");"
data = tuple([vid,sid]+sampledatalist+[variant_allele_count,alleles_genotyped])
c.execute(sql,data)
if(linecounter%10000 == 0):
print "processed " + str(linecounter) + " rows."
linecounter += 1
# do some indexing to improve performance
sql = """
create index var_chrom_idx on vartest.variants(chrom);
create index var_pos_idx on vartest.variants(pos);
create index var_id_idx on vartest.variants(id);
create index eff_vid_idx on vartest.effects(vid);
create index eff_effect_idx on vartest.effects(effect);
create index sv_vid_idx on vartest.sample_variants(vid);
create index sv_sid_idx on vartest.sample_variants(sid);
"""
c.execute(sql)
# close everything
infile.close()
c.close()
conn.close()
Running time: 4 hours. update 2012-10-01: I also found it useful to create a table summarizing the information in the sample_variants table:
create table sample_variant_summary as
select sv.vid, min(gq::numeric) min_gq, avg(gq::numeric) av_gq, min(dp::numeric) min_dp, avg(dp::numeric) av_dp,
sum(sv.alleles_genotyped) alleles_genotyped,
sum(case when s.phenotype = 1 then variant_allele_count else 0 end) case_alleles,
sum(case when s.phenotype = 0 then variant_allele_count else 0 end) control_alleles,
sum(case when s.phenotype = 1 and variant_allele_count = 2 then 1 else 0 end) case_hom,
sum(case when s.phenotype = 0 and variant_allele_count = 2 then 1 else 0 end) control_hom,
sum(case when s.phenotype = 1 and variant_allele_count = 1 then 1 else 0 end) case_het,
sum(case when s.phenotype = 0 and variant_allele_count = 1 then 1 else 0 end) control_het
from sample_variants sv, samples s
where sv.sid = s.sample_id
and gq <> 'NA' and dp <> 'NA'
group by sv.vid
order by sv.vid
;
create index svs_vid_idx on sample_variant_summary(vid);
11. converting to plink format: vcftools PLINK can do a whole GWAS for you, if your needs are pretty standard. But even if you’re planning to do your own modeling and analysis, you might want to use PLINK to model population substructure, which is to say, to group samples into different groups (ethnic groups or families) in order to control for their relation to each other. To get started with PLINK, you have to have ped and map files, or the transposed version, which are tped and tfam files. GATK a VariantsToBinaryPed tool, but as of today it has some unresolved issues with reading sample metadata. Until these are resolved, it’s simple enough to use vcftoolsinstead:
vcftools --vcf variants.vcf --plink-tped
This took about 30 seconds. Instead of using the out.tfamfile that vcftools creates, I created my own since I already have my samples’ metadata in PostgreSQL:
copy (
select sample_id, sample_id, 0, 0, sex, phenotype from samples
) to 'c:/your/path/here/variants.tfam';
If you need to create a samples table first, be sure to use PLINK’s coding for sex and phenotype. With that, you’re ready to run PLINK. A sample call to which common (MAF> 5%) variants are most highly associated with your phenotype:
plink --tfile variants --no-parents --1 --maf .05 --fisher --adjust --out just_an_example
If you haven’t used it, the PLINK tutorial is a great primer. update 2012-10-23: now that you’re done with calling variants, you’ll probably want to QC those as well. I provide some instructions in this post. end update conclusions In terms of getting oriented and learning how to use the tools, I found that the overall startup cost of figuring out GATK was higher than for the tools in the bowtie2/samtools pipeline I’ve previously posted. Key to getting this pipeline to work this time was basing as many steps as possible on tools, pipelines and resources provided “in house” by Broad for use with GATK, which helped me avoid the compatibility issues I had been having earlier. On the plus side, Broad has a team of responsive, helpful staff devoted to GATK, and has created a well-organized community resource for GATK, the likes of which I haven’t seen for any other tool in this domain. In terms of quality of output, I got better results with this GATK pipeline (for instance, my transition/transversion ratio looks more like what I expected). I also got higher coverage of target (~44x) with GATK/BWA than with bowtie2 (~40x), indicating that more reads were aligned successfully, but of course there’s no fair comparison here since this pipeline uses a 20+hour sensitive alignment whereas I ran my other pipeline in --very-fast mode for 3 hours. I haven’t done the kind of test that would allow me to directly compare the two. A note on reference genomes: I ran this whole pipeline using hg19 and it worked out all right for me, but after learning that snpEff discourages the use of hg19 I started to think that perhaps GRCh37 would have been better from the beginning (even though the two are basically the same). A final thought: I came back to GATK after trying other tools because it seems to be increasingly the gold standard in its realm. While I haven’t done enough comparing to be able to enumerate the dimensions on which it outperforms its peers, the fact that it is well-respected and widely used is a benefit in and of itself, as you’ll have less to explain to your colleagues, reviewers, or anyone else who sees the analysis you do with your resulting data. Comparability with existing datasets is also important. For instance, if you want to combine your samples with 1000 genomes samples in order to calculate IBD using a larger pool, you’ll be glad you used GATK because the 1000 genomes project also uses GATK. Finally it seems that there is a steady stream of improvements to GATK coming down the line from Broad so new features, fixes and improved accuracy are all in the works.
