Making sense of the Allen Institute mouse brain images
In a meeting earlier this week I was finally able to get clarification on what all I’m looking at in the Allen Institute images mentioned in a previous post.
ISH images
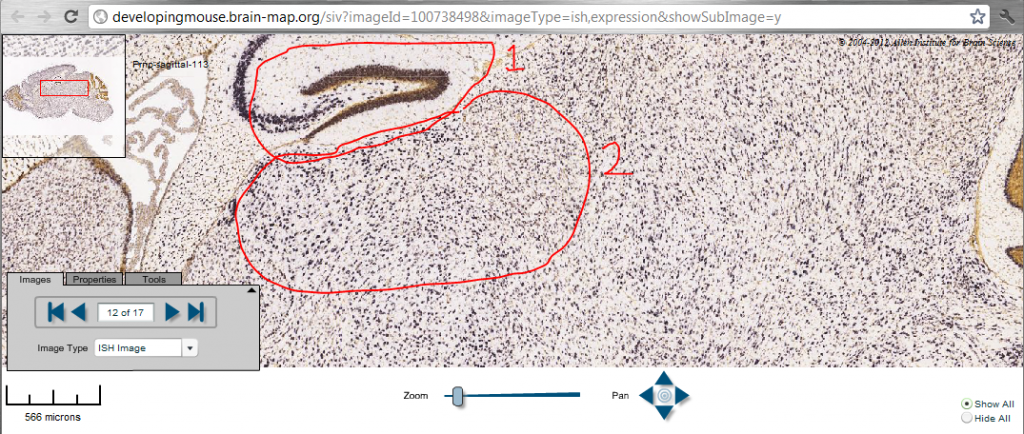
In these images, purkinje cells are stained purple, while granular cells are stained brown with hematoxylin (aka HE or H&E). Purkinje cells do not stain well with HE. The white regions represent the “molecular layer”. Note that in this series of sagittal sections, 17 is the midline and 1 is the outermost section, so 10-12 is around the region where damage to the thalamus from FFI would be most visible. In the image below, (1) is the “arrow” characteristic of the hippocampus; the part of the thalamus immediately below it, labeled as (2), is the most affected in FFI.
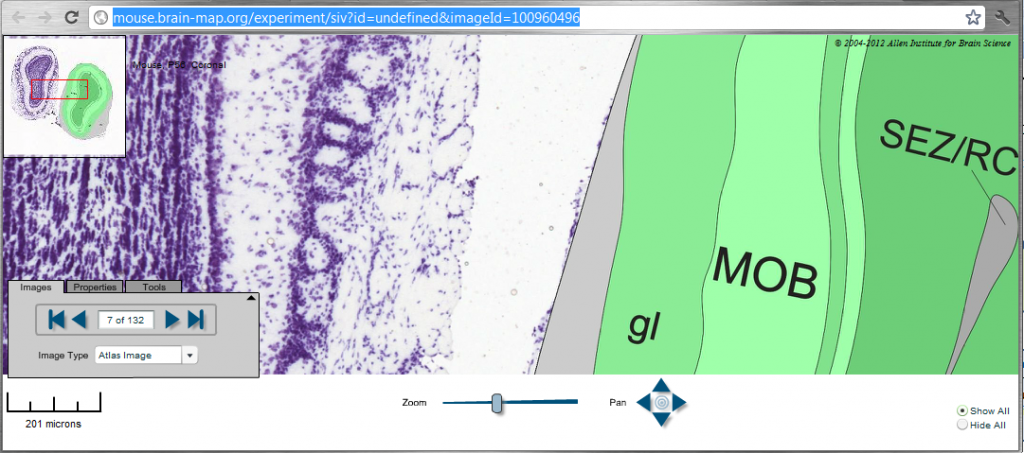
This series shows mRNA expression of a gene of your choice (PRNP in the example below) via a DIG stain where nitroblue tetrazoleum brings out a blue precipitate, and Tyramide Signal Amplification is used to read the image. The images are paired with maps of the brain region you’re looking at. These could also be used towards machine learning for identifying brain regions. But mRNA expression won’t be especially useful for studying FFI because for PRNP Ford et al (2002) have found that mRNA and protein expression are uncorrelated.
http://mouse.brain-map.org/
Connectivity
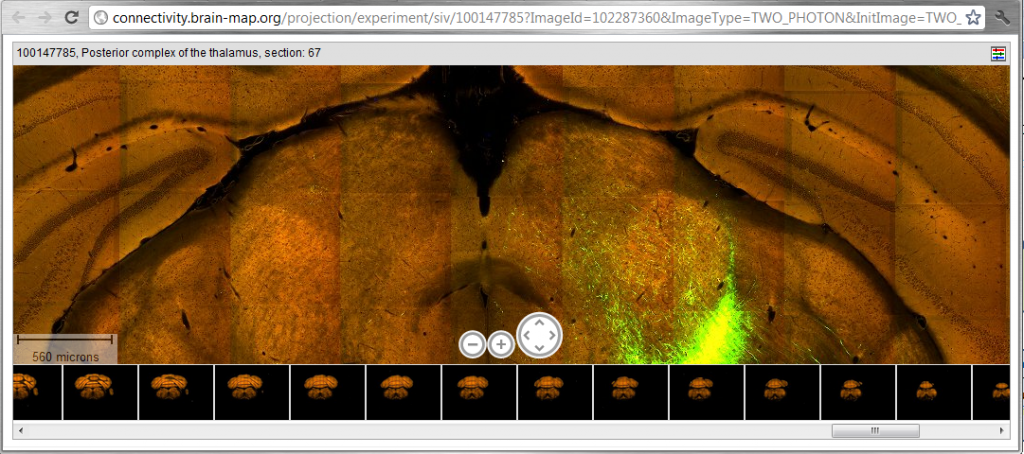
This series shows neuronal connections which will eventually be quite useful to us to figure out what the thalamus connects to: