Regulation of prion protein transcription
Recently I’ve become interested in depleting PrP as a strategy for treating prion diseases. There are several different places to try to intervene in PrP’s life cycle, the earliest of which is transcription. Therefore this post aims to summarize what is known about how PrP’s transcription is regulated, and to assess whether there are opportunities for therapeutic intervention to disrupt transcription of PrP.
defining the prion promoter
The first effort to characterize the promoter region of PrP concluded that a 273bp region was necessary for regulating PrP transcription [Mahal 2001]. Mahal undertook a series of deletion experiments, transfecting HeLa cells with plasmids that expressed luciferase under various deletion mutants of the sequence near the PrP transcription start site (TSS), in order to identify the crucial region without which luciferase was not expressed. The resulting 273bp region was defined as “-148 to +125, relative to the cap site”. ”Cap” appears to refer to the 5′ cap of the PRNP transcript, i.e. the transcription start site. In human genome hg19 coordinates, PRNP RefSeq transcript variant 1 is transcribed from chr20:4666797-4682234. (Note the RefSeq link gives length and sequence for the mRNA, while the UCSC link shows the entire pre-mRNA.) If chr20:4666797 were indeed the TSS to which Mahal referred, then the promoter would extend from chr20:4666649-4666922, but when I get DNA for that region it doesn’t quite match Mahal’s DNA, so Mahal must have been working from a different transcription start site. Instead, the 273bp promoter identified by Mahal appears to be located at precisely chr20:4,666,877-4,667,149. Here’s the sequence:
>hg19_dna range=chr20:4666877-4667150 5'pad=0 3'pad=0 strand=+ repeatMasking=none CAAGCGAATCTCAACTCGTTTTTTCCGGTGACTCATTCCCGGCCCTGCTT GGCAGCGCTGCACCCTTTAACTTAAACCTCGGCCGGCCGCCCGCCGGGGG CACAGAGTGTGCGCCGGGCCGCGCGGCAATTGGTCCCCGCGCCGACCTCC GCCCGCGAGCGCCGCCGCTTCCCTTCCCCGCCCCGCGTCCCTCCCCCTCG GCCCCGCGCGTCGCCTGTCCTCCGAGCCAGTCGCTGACAGCCGCGGCGCC GCGAGCTTCTCCTCTCCTCACGAC
As far as I can tell, the term ‘prion promoter’ is used loosely, and may refer to exactly this 273bp, or to a larger or smaller region. Borchelt 1996 pioneered the use of a much larger swath of PrP regulatory elements as a promoter to drive expression of transgenes in mouse models of other human diseases (unrelated to PrP), for instance the N171-82Q Huntington’s Disease mice. And Asante 2002 reported using a yet smaller, 214bp prion promoter to drive transgenes. Transcriptional regulation is complex: regulatory elements in DNA can exist far upstream or downstream of the transcription start site, and we don’t always understand how they play their regulatory role, so it’s unlikely that Mahal’s critical region is the only region that matters.
Mahal’s bioinformatic analysis predicted binding sites in the prion promoter for Sp1, a transcription factor involved in embryonic development, along with AP-1 and AP-2. But as far as I can tell, no one actually tried to determine experimentally which transcription factors bind, or how PrP transcription is regulated, until a few years later.
stress response
The earliest study I could find that addressed PrP transcription [Shyu 2002] found that PrP protein was upregulated in response to heat shock (cells placed at 42 C). PrP mRNA was never measured. Instead, a reporter construct with luciferase under the prion promoter was used to confirm transcription as the mechanism of upregulation. A predicted binding site for heat shock transcription factor 1 (HSF1) was found in the promoter, and when this binding site was deleted, the luciferase response was greatly reduced. Shyu then looked for HSF1 binding to the prion promoter using an electrophoretic mobility shift assay (EMSA) [reviewed in Hellman & Fried 2007], and confirmed binding.
Years later, Steele 2008 (ft) infected HSF1 knockout mice with prions and found that they succumbed to prion disease about 20% sooner than wild-type mice. Curiously, they did not have elevated PrPSc production nor infectivity nor earlier onset of symptoms – only fatality was accelerated, implying perhaps a reduced ability to cope with the PrPSc. In any event, Steele found no difference in baseline PrP levels in the brains of the HSF1 knockout mice vs. wild-type. That doesn’t rule out a role for HSF1 in regulating PrP under heat shock conditions, but it does suggest HSF1 is not important for PrP transcription under normal conditions in vivo.
Another study [Wang 2005] found slightly elevated PrP protein and mRNA following exposure of N2a cells to oxidative stress agent nitric oxide (NO) and the inflammatory agent lipopolysaccharide (LPS). NO was hypothesized to act through a guanlyl cyclase → MEK → p38 MAPK pathway, and pharmacological inhibitors of these proteins reduced PrP levels. The authors were not super rigorous about proving that this pathway regulates PrP: there was no knockdown and rescue, no confirmation of DNA binding, and many of the effects were of marginal statistical significance. Another study reported that pharmacological MEK inhibitors abolished PrP-res, in cell culture [Nordstrom 2005 (ft)]. In Nordstrom’s Fig 3 you can see that both PrP-res and total PrP are reduced after treatment. But there was no deeper investigation as to whether the MEK inhibitors acted through a transcriptional mechanism.
Another study hypothesized that the unfolded protein response (UPR) transcription factor XBP-1 might be involved in prion pathogenesis, but found no difference in disease course nor PrP levels in XBP-1 knockout mice [Hetz 2008]. A later study suggested that, at least in breast cancer, ER stress does induce PrP transcription via XBP-1 and BiP [Dery 2013 (ft)]. PrP levels are elevated in several cancers, and high PrP levels are associated with a poorer prognosis, reportedly because PrP promotes cell survival. Dery found that ER stress-inducing agents caused an increase in XBP-1 mRNA and prion protein levels, and one experiment with siRNA against XBP-1 reduced PrP levels. Mutation of predicted ER stress response element (ERSE) sequences in the prion promoter reduced the PrP transcriptional response.
copper response
It’s been known for a while now that PrPC binds Cu2+ ions at its PHGGGWGQ octapeptide repeats [Brown 1997, Stockel 1998] and that Cu2+ binding causes PrPC endocytosis [Pauly & Harris 1998].
It was later shown that exposing primary cultured neurons to copper causes an increase in PrP mRNA and protein [Varela-Nallar 2006 (ft)]. This led to a natural hypothesis as to what proteins might be regulating transcription. A class of proteins called metallothioneins, which manage heavy metals, are regulated by metal transcription factor 1 (MTF-1) [Heuchel 1994 (ft)], which binds to a a sort of loosely defined DNA sequence motif called a metal response element (MRE) [Giedroc 2001]. Varela-Nallar describes the MRE motif as “a highly conserved 7-base pair functional core motif [5′-TGC(A/G)CNC] flanked by a less conserved 5-base pair GC-rich domain”. Varela-Nallar identified a few sites that almost matched the MRE motif, located ~2000-2500 bases upstream of the PRNP TSS, and through deletion experiments (similar to Mahal’s method) found that one of these MREs (“−2,653 TGCGtCCCCTGC”; I was unable to find this sequence near PRNP in hg19), as well as some nearby non-MRE regions, appeared to be pretty important for copper-induced PrP expression. Varela-Nallar then looked for MTF-1 binding to that particular 12bp sequence using an electrophoretic mobility shift assay (EMSA) [reviewed in Hellman & Fried 2007]. The EMSA did not reveal any MTF-1 binding to this particular 12bp sequence; also, zinc tends to activate MTF-1 yet had no effect on PrP levels. For both of these reasons, Varela-Nallar concluded that MTF-1 was not involved in the transcriptional regulation of PrP.
A more recent study has disagreed [Bellingham 2009 (ft)]. Bellingham’s study relied on cultured fibroblasts from a patient with Menkes disease. Menkes disease is an X-linked disease caused by loss-of-function mutations in the gene ATP7A, which encodes Menkes protein, a P-type ATPase which pumps copper out of the cytosol and into the secretory pathway [Barnes 2005 (ft)]. Patients with Menkes disease accumulate excess intracellular copper. Bellingham compared the original unaltered patient fibroblasts, dubbed MNK(Del), to ones transfected with a vector overexpressing ATP7A, dubbed MNK(++), as well as an empty vector control, MNK(v/o). The comparison of these lines was striking: PrP expression appeared to be completely abolished in the MNK(++) lines – neither the protein nor mRNA were detectable at all (Fig 3). Bellingham hypothesized that Sp1 and MTF-1 were involved in copper-dependent regulation of PrP and sought to confirm this experimentally. In the MNK(Del) cells, knocking down Sp1, MTF-1 or both reduced PrP levels, and transfecting the cells with additional Sp1, MTF-1 or both increased PrP levels. The MNK(++) cells stubbornly refused to express any PrP at all under any conditions studied.
The most striking aspect of Bellingham’s study is the suggestion that copper is absolutely necessary for PrP transcription – at least in fibroblasts, under the conditions studied, etc. If it’s true, it begs the question of what transcription factors mediate this copper dependence. MTF-1 is well known to respond to copper [Heuchel 1994 (ft)], and interestingly, it appears to be on call for both copper surplus and copper deficit, which it manages through transcription of different genes, at least in Drosophila [Selvaraj 2005 (ft)]. Sp1 has also been reported to respond to copper [Song 2008 (ft)]. Bellingham didn’t do an EMSA or any other experiments to confirm that either of these proteins actually binds to any location in the prion promoter. However, the finding that overexpression and knockdown both affected PrP levels is compelling functional evidence that they’re involved in some way, and seems to outweigh the earlier claim that MTF-1 is not involved, which was based on a lack of binding to just one predicted site [Varela-Nallar 2006 (ft)].
Another study reported a different copper-dependent transcriptional mechanism for PrP [Qin 2009 (ft)]. Qin examined the time course of PrP mRNA and protein elevation in response to Cu2+ in some detail and concluded, as these other authors have, that the PrP response to copper is transcriptional in nature. Qin proposed a pathway whereby copper causes phosphorylation of ATM at S1981, and activated ATM then activates Sp1 through the MEK/ERK pathway, and activates p53 by phosphorylation at S15. Qin provides some pretty rigorous evidence for this in Fig 4, by using siRNAs to knock down various elements of the pathway and show that this abolishes the response in all the downstream elements. Importantly, Qin also used an EMSA to show direct binding of both Sp1 and p53 to the prion promoter. Qin proposes that the upregulation of PrP in response to copper may constitute a stress response, since copper causes the creation of reactive oxygen species.
Qin’s finding that the MEK/ERK pathway is involved in PrP transcription has been subsequently validated and extended [Cisse 2011 (ft)]. Cisse found that depletion of ERK1 or dominant negative inhibition of ERK1 or MEK reduced PrP mRNA levels, and transfection of cells with ERK1 or MEK increased PrP transcription. Cisse notes that ERK1 activation can promote gene transcription either by phosphorylation of Sp1 or by phosphorylation of c-Fos, part of the AP-1 complex. Mutation of the predicted AP-1 binding sites also reduced PrP transcription, but mutation of the predicted Sp1 binding sites had no effect, suggesting AP-1 was responsible. Interestingly, ERK1 also controls PrP proteolytic cleavage by ADAM10 [reviewed in Checler 2012].
amyloid intracellular domain
Here’s a lightning-quick review of the metabolism of amyloid precursor protein (APP). Beta-secretase (BACE1) cleaves it on the exoplasmic side, and gamma-secretase (including PSEN1 & PSEN2) cleaves it on the cytosolic side, producing an extracellular fragment amyloid beta (Aβ) with roles in Alzheimer’s disease, and an intracellular fragment called the amyloid intracellular domain (AICD) [Vassar 2009].
PrP and amyloid beta interact physically, leading to downstream signaling changes that are only now beginning to be understood – see my latest PrP/Aβ post. PrP also regulates the production of Aβ through its inhibition of BACE1 [Parkin 2007, Rushworth 2013]. It turns out that APP metabolism may also regulate PrP: it has been reported that the AICD activates transcription of PrP [Vincent 2009 (ft)].
Vincent found that knockout of PSEN1 and PSEN2 in fibroblasts reduced PrP mRNA and protein by about half. betaAPP overexpression increased PrP, betaAPP knockout reduced PrP, and direct expression of AICD fragments C50 or C59 increased PrP. Vincent proposes that p53 mediates the AICD’s effects on PrP. p53 knockout reduced PrP expression (rescued by p53 cDNA transfection) and abolished the stimulatory effects of C50 and C59; ChIP confirmed direct binding of p53 to the prion promoter. Vincent’s group had previously reported that AICD activates transcription of p53 [Alves da Costa 2006], so this is postulated as the mechanism for the AICD → p53 → PrP connection. update 2013-11-04: Vicki Lewis et al failed to replicate the AICD connection – see comment.
no unbiased screens so far
All of these studies started from a particular hypothesis about what regulates PrP transcription and then sought to prove it. Most started from completely different hypotheses and did not explore or directly seek to confirm one another’s findings.
No study has yet utilized an unbiased screen for proteins binding to the prion promoter. One unbiased method for identifying regulators of a particular promoter is to prepare a biotinylated synthetic DNA oligomer of the desired promoter, dip it into a nuclear extract, and then perform mass spectrometry on the proteins that come out bound to the DNA [Nordhoff 1999]. This method has its pitfalls – it by no means captures the range of physiological conditions under which transcription might be stimulated – but at least it is unbiased and has the potential to catch things that no one has thought to hypothesize.
Something big and unbiased has happened since 2009, though: the ENCODE project.
ENCODE data
ENCODE (Encyclopedia of DNA regulatory elements) aims to create a genome-wide map of everything that regulates gene expression: histone modifications, DNA methylation, and transcription factor binding among other things. Most of the data comes from hundreds of genome-wide ChIP-seq experiments. DNA-binding proteins are cross-linked to DNA with formaldehyde, one protein of interest is pulled down with antibodies, de-cross-linked, and then the DNA that had been attached to it is sequenced. This gives you a map of where in the genome the protein had been bound.
Though ENCODE is unbiased in that it is genome-wide, it is limited to the transcription factors that were chosen to be studied. The full list of ChIP-seq experiments includes Sp1 but neither p53 nor MTF-1. ENCODE currently has transcription factor binding data from 690 experiments on 161 transcription factors, but finding the data for all of them is confusing. If you simply turn on the ENCODE Txn factor track in the UCSC genome browser, you’ll see only 26 of the 690. If you instead visit the Uniform TFBS Track page you’ll be confronted with the entire list of 690 and no way to display data from all of them except by checking each and every box.
The easiest way I’ve found of getting the data for all 161 transcription factors is through the UCSC table browser, settings as shown below. I used chr20:4665648-4668377, so ~2700bp, a 10x zoom-out from Mahal’s promoter region.
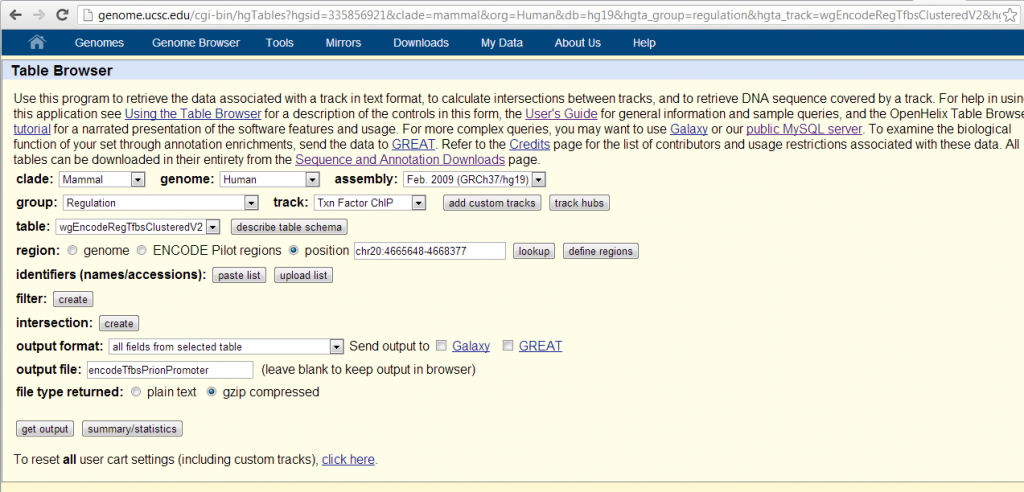
Results are here: [TXT]. The table contains 48 entries, apparently only the transcription factors with scores > 0. Three transcription factors got the maximum possible score of 1000: JunD, FOSL2 and c-Fos. All three of these are components of the AP-1 transcription factor complex, thus confirming Mahal’s original bioinformatic prediction of AP-1 binding [Mahal 2001]. The thickStart and thickEnd columns tell you where exactly the peaks are; FOSL2 and c-Fos both overlap Mahal’s 273bp promoter region of chr20:4666877-4667149. Besides the AP-1 proteins, STAT3 also comes close to the maximum, with a score of 950. Sp1 is not on the list, implying it had a score of 0. Some AP-2 transcription factors are present lower on the list.
summary
This table attempts to summarize the findings from the above studies.
| Tx factor | functional evidence | physical evidence | proposed regulatory pathways |
|---|---|---|---|
| p53 | Knockdown [Qin 2009 (ft),Vincent 2009 (ft)], overexpression [Qin 2009 (ft)] | EMSA [Qin 2009 (ft)], ChIP [Vincent 2009 (ft)] | PSEN1&2+APP→AICD [Vincent 2009 (ft)], copper-induced via ATM→MEK→ERK pathway [Qin] |
| Sp1 | Knockdown [Qin 2009 (ft), Bellingham 2009 (ft)], overexpression [Bellingham 2009 (ft)] | EMSA [Qin 2009 (ft)] | Copper-induced via ATM→MEK→ERK [Qin 2009 (ft)] or unknown mechanism [Bellingham 2009 (ft)] |
| AP-1 | Mutation of predicted binding sites [Cisse 2011 (ft)], ChIP-seq [ENCODE] | Activated by ERK1 [Cisse 2011 (ft)] | |
| MTF-1 | Knockdown, overexpression [Bellingham 2009 (ft)] | Copper-induced [Bellingham 2009 (ft)] | |
| XBP-1 | Knockdown, overexpression [Dery 2013(ft)] | Mutation of predicted binding sites [Dery 2013(ft)] | ER stress [Dery 2013(ft)] |
| HSF1 | Luciferase reporter [Shyu 2002] | Deletion of predicted binding site, EMSA [Shyu 2002] | Heat shock response [Shyu 2002] |
If we very generously (i.e. unskeptically) believe everything that’s been reported, the picture of prion protein transcription that has been developed so far looks as follows:
If we’re more skeptical and only consider those things which (A) are supported by more than one study and (B) for which both physical and functional evidence are available, the picture thins considerably:
In either case, what we have at this point is certainly a very incomplete picture of how PrP transcription is regulated. Most of these studies focused on transcriptional upregulation in response to a particular environmental stress, rather than characterizing what drives baseline transcription levels. Only recently [Cisse 2011 (ft)] was AP-1 activity implicated in the literature, even though according to ENCODE it is the most active transcription factor in the prion promoter.
The most important piece that is missing is what drives neuronal transcription of PrP. PrP mRNA is expressed ~10 times higher in the CNS than in most peripheral tissues [Novartis BioGPS microarray data], implying there must be some neuronal-specific transcription factors, upstream regulatory mechanisms or epigenetic marks at work. Of the above-cited studies, just one used primary cultured neurons [Varela-Nallar 2006 (ft)] and two used CNS-derived cancer cell lines (N2a and NT-2) [Qin 2009 (ft), Shyu 2002 respectively]. The rest used fibroblasts or peripheral cancer cell lines. Browsing the ENCODE human cell experiment list, I also don’t see any CNS cell types there.
assessment of therapeutic potential
It’s unclear if any of these pathways lend themselves to therapeutic intervention. The most potentially interesting finding is that cytosolic copper depletion by overexpression of ATP7A reduces PrP expression to undetectable levels [Bellingham 2009 (ft)]. If this result (originally from fibroblasts) could be replicated in neurons, and could then be associated with the inactivation of one particular transcription factor or one particular protein upstream thereof, that could potentially constitute a drug target. If Bellingham is right that MTF-1 is involved in some way, it is interesting to note that MTF-1 knockout is embryonic lethal, perhaps due to liver decay [Gunes 1998 (ft)] but that Cre-mediated adult knockout in the liver is survivable [Wang 2004 (ft)]. As far as I can tell from the literature no one has done a Cre knockout of MTF-1 in neurons, and no one has checked whether PrP is expressed in MTF-1 knockout cells.
Transcription factors are not the ideal drug targets – they tend to promote hundreds of genes, so inhibiting them to turn off one substrate is truly razing a village to catch one man. But blocking a transcription factor is not entirely unheard of – for example disulfiram inhibits NF-KB [Schreck 1992 (ft)] and has been explored for treatment of HIV and cancer, though its relevant mechanism of action in those diseases may be something different [Lovborg 2006 (ft), Doyon 2013]. Also, disrupting a particular DNA binding event or a recruitment interaction between two transcription factors can be more specific than just inhibiting a transcription factor altogether.
