How prion protein was first purified
Months ago, I ran across the original series of papers in which Stanley Prusiner achieved a homogeneous purification of prion protein and began to decipher its amino acid sequence [Prusiner 1982a, Prusiner 1983, Prusiner 1984]. This purification process seemed like quite a feat, but I couldn’t begin to understand the methods used. After attending a biochem lecture a couple of weeks ago covering methods for protein purification, I thought I was finally equipped to understand this, so I went back and took a deeper look at these old papers.
The process of purifying PrP was amazingly involved, and the road to get there even moreso. Prusiner spent much of the 1970s fractioning hamster brain homogenate on every imaginable dimension and assaying for which fraction contained infectivity, a journey partly chronicled in the seminal “prion” paper [Prusiner 1982b]. The purification procedure that finally emerged from this effort is presented in [Prusiner 1983] and is nicely summarized in [Prusiner 1984] as well.
In Step 1, the starting material – scrapie-infected brain homogenate – is simply ‘clarified’, which I understand to mean that large particles and impurities are removed. This is accomplished with continuous flow centrifugation, which Eppendorf describes as being for “solids removal”. A liquid is pushed up into the bottom of a rapidly rotating tube, and solids are retained on the walls of the tube while liquid is forced out the top. The centrifugation speed is moderate (compared to later steps) at just 10,000 rpm. As in any centrifugation process, the solids are referred to as the pellet (P1 since this is Step 1) and the liquid is the supernatant (S1).
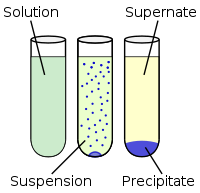
In Step 2, PrPSc is precipitated out based on its property of being detergent-insoluble. The liquid from the previous step, S1, is mixed with two detergents: Triton X-100 and sodium deoxycholate, and then precipitated with PEG-8000 as a precipitant. Precipitation means causing a solute (in this case PrPSc) to form particles in suspension, which can then be centrifuged into a pellet, as shown in these three steps:
(After reading about polyethylene glycol, of which PEG-8000 is simply the term for polymers of ~8,000 kDa molecular weight, I still don’t understand what makes it act as a precipitant.) The precipitated material was pelleted out, again using continuous flow centrifugation but this time at a higher speed, 16,000 rpm.
In Step 3, the pellet from Step 2 (P2) is re-suspended in a solution with a buffer (Tris OAc), a detergent (Triton X-100), a reducing agent (DTT), and a salt (CaCl2), none of whose functions in this context I understand. The suspension is then digested with micrococcal nuclease, which breaks down RNA and DNA – an important step for removing contaminating nucleic acids and, more importantly, for proving that the “scrapie agent” didn’t require nucleic acid.
The suspension was then also subjected to a light proteinase K (PK) digestion. PK, mind you, can abolish scrapie infectivity – one early piece of evidence for the prion hypothesis – but its activity is dependent on enzyme concentration (more is more), time (more is more), and temperature (most active at 37°C). The digestion in this protocol was 8h of 100 ug/mL of enzyme at 4°C. That’s not much – the protocol required to denature 99.9% of infectivity is more like 30h of 500 ug/mL at 37°C [Prusiner 1981]. So I think my understanding is that this light PK digestion is here mostly to degrade other proteins, thus comparatively enriching for the at least partially PK-resistant desired protein: PrPSc.
After the two digestion steps, the suspension was then again treated with two detergents (Sarkosyl and sodium deoxycholate) and precipitated (with ammonium sulfate) and then centrifuged again (now at 19,000 rpm) to yield P3.
In Step 4, P3 is again re-suspended and then subjected to zonal rotor centrifugation to separate sucrose gradients. ”Zonal rotor” just means that instead of tubes full of liquid inside a centrifuge, the whole centrifuge is directly full of a liquid. The principle of sucrose gradients is quite interesting: the centrifuge is filled with distinct layers or a continuous gradient representing different concentrations of sucrose – in this case, a continuous gradient from 60% (at the outside) to 0% (on the inside). Molecules will “sediment” according to density – a molecule with the same density as a 40% sucrose solution will settle in the 40% fraction. Sucrose gradient layers are often referred to in “Svedbergs” (e.g. “the 26S proteasome”) though this is technically a measure of time rather than of density or sucrose concentration.
Prusiner assayed every different fraction of this gradient for scrapie infectivity, and found that infectivity peaked somewhere in the range of about 30-40% sucrose [Prusiner 1983, Fig 1].
This was enough purification to begin to study prions at the molecular level in ways that hadn’t been possible. Prusiner performed electron microscopy on the most infectious sucrose gradients and saw 100-200nm long rods of aggregated PrPSc, which resembled amyloid fibrils from Alzheimer’s brains [Prusiner 1983, Fig 4]. He also boiled the fractions in another detergent (sodium dodecyl sulfate, SDS), which inactivated infectivity but also disaggregated the rods, thus making it possible to study pure PrP monomers for the first time. He ran the resulting monomers through SDS-PAGE, a process whose principles I only recently came to understand. If you just put proteins on a polyacrylamide gel and run an electrophoresis (PAGE), they’ll separate according to charge, with cationic proteins headed towards the anode and anionic proteins bound for the cathode. By first treating proteins with SDS, you uniformly coat them in negative charge so that they all migrate towards the cathode, distinguished only by their molecular mass – extra mass hinders their progress. So this is a method for determining the molecular mass of an isolated protein. Mind you, this looks just like a Western blot [Prusiner 1983, Fig 2] but a Western blot is an SDS-PAGEd protein transferred to nitrocelluose and then blotted with antibodies. In 1983 the world had never seen no antibodies to PrP yet because prion diseases don’t cause an immune response and the protein had never before been purified. Instead, the protein had to be labeled using radioactive iodine and Bolton-Hunter reagent and then developed as an autoradiogram.
The next year, Prusiner was back with another huge breakthrough: the beginnings of a characterization of the amino acid sequence of PrP [Prusiner 1984]. This study used the same purification protocol as before, plus an additional step of a proteinase K digestion (4 ug/mL at 37°C for 2h), apparently necessary to disaggregate the rods enough to make it possible to hydrolyze (break down) the peptide bonds between every amino acid. Subjecting PrP to protease digestion also meant that only the protease-resistant fragment was being studied, a fragment dubbed PrP 27-30. Even after the protein was run through high-performance liquid chromatography (HPLC) to isolate it based on molecular weight, its mass still ranged from 27 to 30 kDa, probably reflecting differences in glycosylation, beta cleavage site and/or PK-resistant fragment size.
To figure out the amino acid composition (not sequence) of PrP, Prusiner digested all of the peptide bonds and then ran the free amino acids through high-performance liquid chromatography (HPLC), separating the amino acids by mass. At least at that time, this technique was unable to distinguish D from N and E from Q, but it did show that PrP contained at least 17 of the 20 amino acids (in fact, it contains all 20). This disproved what had apparently been one hypothesis about prions up to that time: that perhaps they were able to replicate without nucleic acid because they were just simple repetitive sequences of a couple of amino acids that happened to code for a protein that produces itself.
Prusiner also determined the amino acid sequence of the N-terminal edge of PrP 27-30: GQGGGTHNQWNKPSK, corresponding to codons 81-95 of SHaPrP. Interestingly, weaker signals were observed for sequences starting at other codons, revealing heterogeneity in where the PK-resistant fragment ended.
When I first saw this paper, I couldn’t begin to understand how amino acid sequencing was possible in 1983 considering that, 30 years later, we still don’t have any really high-throughput methods for sequencing novel proteins today. Mass spec proteomics is still horrifyingly expensive and even then, the quantification of proteins involves essentially a lookup table (UniProt) of known proteins – a far cry from the ability of next-gen DNA sequencing to read an arbitrary sequence.
But as I learned recently, amino acid sequencing has actually been around since 1949 , just at very, very low throughput [Edman 1949 (ft)]. Though Prusiner does not go into the technique in detail in the paper, I am assuming it’s basically the same technique, known as Edman degradation. The purified protein is treated with phenylisothiocyanate (“Edman’s reagent”), which (if I understand correctly) reacts with only the exposed N-terminal amino acid of a protein, and makes that amino acid’s peptide bond more vulnerable to cleavage by acids. So: react with phenylisothiocyanate, react with acid, run the one cleaved amino acid through HPLC, wash, repeat. In this manner Prusiner managed to get through 15 amino acids down the N-terminus of the PK-digested protein.
It’s also possible to get a protein’s entire sequence via Edman degradation, by cleaving the protein with different enzymes (e.g. trypsin and chymotrypsin) so that you get different overlapping peptides, separating those by mass, and then sequencing each of those and reassembling the sequences. Prusiner didn’t do this, presumably because the purified protein was too precious – it took 10 liters of a diluted brain homogenate as the starting material to get just a few nanomoles of purified PrP (each aliquot used for a different experiment was 300 – 1000 pmoles). At a molecular mass of ~27-30,000 g/mol, the total quantity probably amounted to something on the order of a few hundred μg of PrP.
It was a battle hard won. The purification and characterization of PrP set the stage for the discovery of PrP mRNA and the revelation that it was encoded in the host genome [Oesch 1985, Chesebro 1985], and for the development of the first anti-PrP antibodies [Bendheim 1984]. All of the tools that make studying prions so easy today – transgenic mouse models, monoclonal antibodies, recombinant PrP – required first being able to study the protein in its pure state.